Human astrovirus (HAstV) 1–8 and highly divergent HAstVMLB1−3 genotypes have been detected in children both with and without acute gastroenteritis (AGE). One hundred and seventy fecal samples from children (≤5 years old) living in the Amazon region were evaluated for the presence of HAstV1–8, HAstV MLB1−3 and HAstVVA1−3, using an usual RT-PCR protocol and a new protocol with specific primers designed to detect HAstVMLB1−3. HAstVMLB1 and HAstV MLB2, as well as the HAstV3 and 5 genotypes were detected. HAstVMLB1−2 genotype was detected for the first time in Brazil at a frequency of 3.5% (6/170).
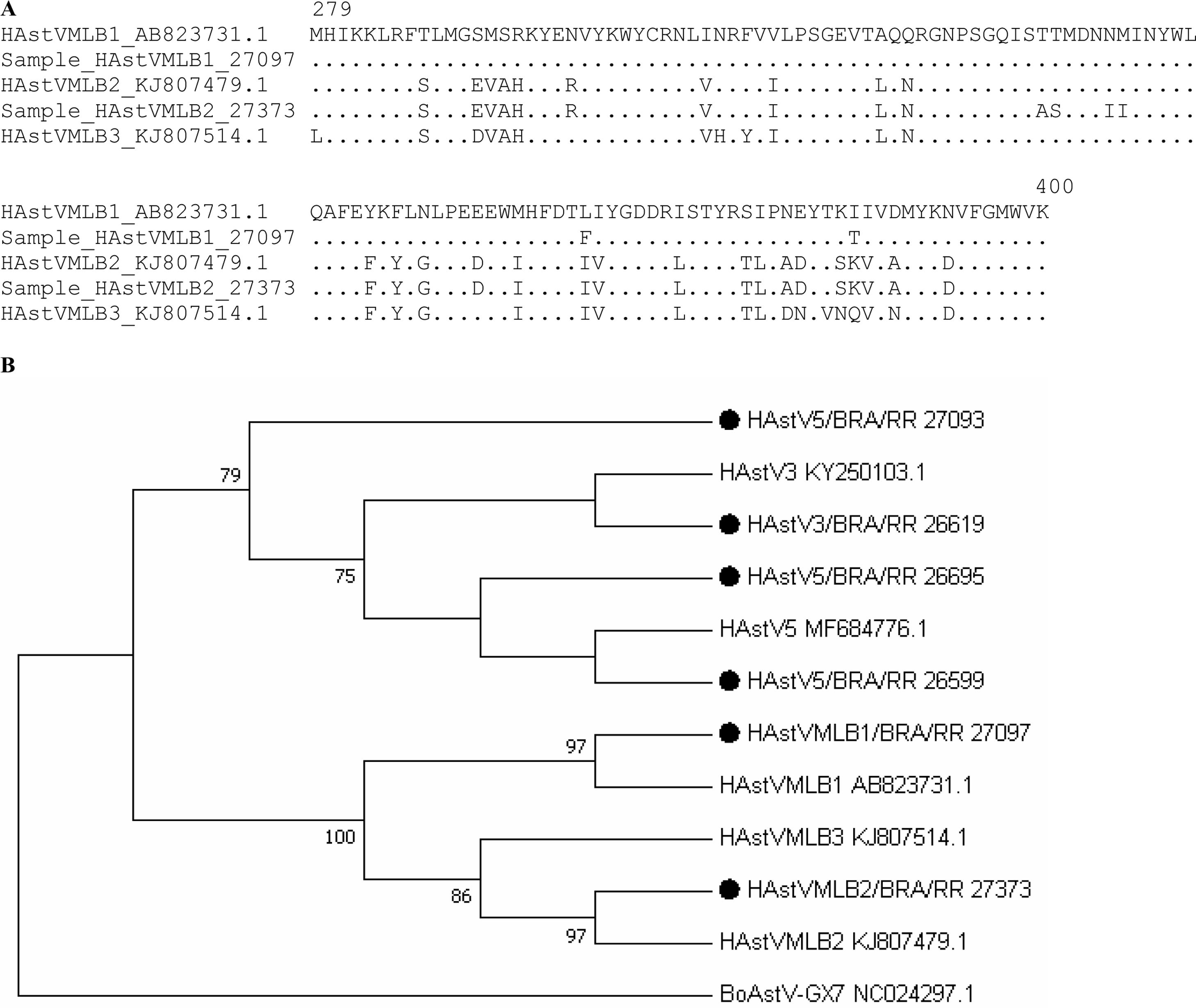
Human astrovirus (HAstV) causes acute gastroenteritis (AGE) in younger children, with a worldwide frequency of 0.5%–15%.1,2 These viruses have a positive sense RNA genome of 6−8 kb in length harboring 5′ and 3′ untranslated regions and three open reading frames (ORFs), namely ORF1a, ORF1b, and ORF2. The ORF1b is translated through a frameshift mechanism and encodes an RNA-dependent RNA polymerase (RdRp).2 The wide host range of the Astroviridae family renders high diversity to these viruses. Two groups Mamastrovirus (MAstV1−19) and Avastrovirus (AAstV1−3) are recognized, based on the phylogenetic analysis of the amino acid sequence (aa) of the full length ORF2 region of the genome that encodes the capsid. MAstV1 and MAstV6 contain, respectively, the HAstV1–8 (“classic”) and the highly divergent (“non-classical”) HAstVMLB1−3 genotypes, both infecting humans. Different lineages of classic HAstV are recognized and have been recently reviewed.2 There is a limited number of capsid sequences available compared to RdRp sequences, and the RdRp has been useful for HAstV detection and genotyping, especially for HAstVMLB1−3.2–4 In this study, the non-classic HAstVMLB1 and HAstVMLB2 as well as the classic HAstV3 and HAstV5 were identified in fecal samples collected from children living in the Amazon region. Previously, no HAstVMLB2 had been detected in Brazil. One hundred and seventy fecal samples, negative for rotavirus A (RVA), GI and GII noroviruses, human adenovirus (HAdV), sapovirus (SaV) (unpublished results) and human bocavirus (HBoV) by quantitative RT-PCR or PCR5 were selected for this study. These samples were collected from October 2016 to October 2017 from children (≤5 years old) receiving care at “Hospital da Criança de Santo Antonio” (HCSA) in the city of Boa Vista, state of Roraima (RR), Northern Brazil. The research protocol was approved in November 23, 2015 by the Ethical Research Committee of Federal University of RR under Number 1.333.480. These children were living in Brazil, Venezuela or Guyana, including demarcated indigenous areas from the Amazon and were hospitalized with AGE. Some of them were Rotarix™ (RV1) RVA vaccinated, as informed by the children’s parents or guardians and verified in the vaccination card. After hospitalization caused by AGE episodes all the children recovered. Complementary study design information as well as collecting, processing and total RNA isolation have been reported previously.5–7 Reverse transcription polymerase chain reaction (RT-PCR) sample screening was carried out initially using an usual and “consensus” protocol with the SF0073/SF0076 (respectively RdRp nucleotide position 3638-3647 and 4042-4024) primers for detection of HAstV1–8, HAstVMLB1−3 and HAstVVA1−3.8 The segment obtained using these primers has a size of 405 bp from the ORF1b (RNA polymerase) region of the AstV genome.8 Thermal cycling conditions have been previously described.8,9 All the samples tested by the consensual protocol were also evaluated using a new RT-PCR protocol named “HAstVMLB1−3 specific”. In this new protocol, the consensus primers were replaced by specific primers these being FMLB2RdRp-Forward (5′AGGACCCCAAACACTACCTAG3′, RdRp nucleotide position 2868-2888) and RMLB2RdRp-Reverse (5′ CATCCCATACAGTGGGACCA3′, RdRp nucleotide position 3082-3063). These primers were designed using Primer Express software v3.0 (Thermo Fisher Scientific, Waltham, Massachusetts, USA) considering the full ORF1b coding the RdRp of classic and non-classic HAstV and a genetic alignment analysis was performed comparing all classic HAstV1–8, non-classic HAstVMLB1−3 and HAstVVA1−3 sequences available in GenBank up to August 2018. The SuperScript III One Step RT-PCR System with Platinum Taq High Fidelity DNA Polymerase® (Thermo Fisher Scientific, Waltham, Massachusetts, USA) was used according to the manufacturer’s recommendations and optimized thermal cycling conditions. HAstV1−6, 8 and HAstVMLB1 sample genotypes from the Regional Rotavirus Reference Laboratory (RRRL-LVCA, Oswaldo Cruz Institute, Fiocruz) were included in all RT-PCR reactions as positive controls in the “consensus” protocol or positive (HAstVMLB1) and negative (HAstV1−6, 8) in the “HAstVMLB1−3 specific” protocol.9 To verify the sensitivity of the “HAstVMLB1−3 specific” protocol, total RNA from sample number 27373 was quantified by Qubit Fluorometric Quantification (Thermo Fisher Scientific, Waltham, Massachusetts, USA) and tested using 1 μg, 100 ng, 10 ng and 1 ng. Additional RT-PCR amplification was performed to confirm the detection of HAstVMLB2 and HAstV5 using a set of specific primers that targeted the ORF2 region capsid as previously described.10 All amplicons were purified using Wizard® SV Gel and a PCR Clean-Up System kit (Promega, Madison, USA) following the manufacturer’s instructions. The purified amplicons were analyzed by Sanger sequencing using a BigDye® Terminator v3.1 Cycle Sequencing Kit and the ABI Prism 3730 Genetic Analyser® (Applied Biosystems, Foster City, CA, USA) and the primers described above. The nucleotide sequence alignment and phylogenetic tree were constructed using the Mega-Molecular Evolutionary Genetic Analysis Version X software.11,12 Similarity was assessed using the Basic Local Alignment Search Tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Using both RT-PCR protocols (“consensus” and “HAstVMLB1−3 specific”) the HAstV frequency was 5.9% (10/170). Within the HAstV “classic” group, HAstV5 was predominantly detected (three samples) and only one HAstV3 was detected. Four HAstVMLB1 and two HAstVMLB2 were amplified using both RT-PCR protocols described in the study design (frequency of 3.5%; 6/170). HAstV were detected in feces from children living in demarcated indigenous areas,6 except HAstVMLB2 samples (27341 and 27373) and the HAstV3 that were collected from children from Boa Vista, capital of RR. There were more RV1 vaccinated HAstV positive children (seven children) than unvaccinated (three children); otherwise all HAstVMLB1 positive children were RV1 vaccinated; however, a HAstVML2 child was not vaccinated. Out of 10 positive cases, 70% (3/10) were children aged > 1 year but ≤ 2 years. No HAstVMLB3 was detected using the “HAstVMLB1−3 specific” protocol. Table 1 summarizes all the results. The sensitivity of the “HAstVMLB1−3 specific” protocol was evaluated in the context of four concentrations of total RNA extracted from feces according to a routine method,5–7 namely 1 μg, 100 ng, 10 ng and 1 ng. All amounts tested were able to provide HAstVMLB2 amplicons. The “consensus” protocol produced 405bp amplicons and were Sanger sequenced. The amplicons from HAstVMLB1 (samples 26683, 27215 and 27361) positive samples did not render good enough quality by Sanger nucleotide sequencing to be evaluated. The nucleotide (nt) sequences from the 27097 (HAstVMLB1) and 27373 (HAstVMLB2) samples were respectively aligned together with the nt sequences with access numbers AB823731.1 and KJ807479.1. These sequences show 93% (27097) identity with the analyzed samples and 97% (27373), using the Basic Local Alignment Search Tool. Fig. 1A shows the aa sequence alignment from the partial RdRp of samples 27373 and 27097, analyzed with HAstVMLB1 (access number AB823731.1), HAstVMLB2 (access number KJ807479.1), and HAstVMLB3 (access number KJ807514.1) as comparative criteria. Fig. 1B shows the phylogenetic tree constructed based on partial amino acid sequences (121 aa) from the partial RdRp according to what had been used previously,8,9 where HAstVMLB1 (sample 27097), HAstVMLB2 (sample 27373) and all “classic” HAstV samples detected in this study were included. HAstV5 samples 26599 and 26695 showed a common proximity with a patient from China (access number MF684776.1) and the sample 27093 was divergent, although classified by ORF1b and ORF2 (capsid region) sequencing as HAstV5. The HAstV3, HAstVMLB1 and HAstVMLB2 samples are closely related to HAstV from Germany (KY2500103.1), Asia (AB823731.1) and Gambia, Africa (KJ807479.1), respectively. The “HAstVMLB1−3 specific” protocol produced a 215bp amplicon, which was Sanger sequenced. All the previous HAstVMLB1 (three samples) and one HAstVMLB2 (sample number 27373) were detected by the conventional protocol. However, one more HAstVMLB2 sample (sample number 27341) was identified. HAstVMLB1 and HAstVMLB2 showed >98% similarity with the reference samples described above. The similarity between HAstVMLB2 samples 27373 and 27341 was >99.9%, with only one aa difference in the RdRp in sample number 27373 (T334A) absent in sample 27341, considering the reference sequence access number KJ807479.1. The RT-PCR amplification using a set of specific primers targeted the ORF2 capsid region,10 rendering 449bp amplicons from samples 27373, 27341 and 27093. After Sanger nucleotide sequencing and alignment, both samples 27373 and 27341 showed similarity above 98% with the reference sample access number MK327365.1 from China. Sample 27093 was confirmed as HAstV5 as previously described. HAstVMLB1 was first reported in Brazil in 2015 through a surveillance study involving 2913 fecal samples collected from children with AGE under five years old.9 The two HAstVMLB1 positive samples were collected from two one-year old children, in two different Brazilian regions: Maranhão and Rio de Janeiro, northeast and southeast states, respectively. Later, in a study involving 483 fecal samples from children with AGE living in the Amazon region (northern region), no non-classic HAstV was detected.13 In our study using a conventional (“consensus”) RT-PCR protocol, four HAstVMLB1 and one HAstVMLB2 were detected. Several factors may have contributed to a higher detection frequency of HAstV in this study as well as the highly divergent non-classic HAstVMLB1−2, including epidemiological changes and ecotourism that affect populations of vulnerable children living in indigenous regions or outside of urban regions. Indeed, Bitencurt et al.16 reported HAstV frequencies of 4.7% (23/488) occurring in children under five years old living in Acre, Amazon region. However, it seems quite clear that the “HAstVMLB1−3 specific” protocol used is more efficient for the detection of non-classic HAstVMBL1−2, which increased the HAstV frequencies detected in this study. We were not able to confirm that the “HAstVMLB1−3 specific” protocol could detect HAstVMLB3, due to unavailability of HAstVMLB3 positive control. A quantitative RT-PCR (qPCR) protocol could be used to detect HAstVMLB3; however, amplicons from HAstVMLB3 positive samples would be required for Sanger sequencing and fully validating this protocol. In fact, the ORF2 amplicons from samples 27373 and 27341 were hard to obtain and RT-PCR conditions had to be improved (data not shown). The primers used for “conventional” and “HAstVMLB1−3 specific” protocols were designed based on the nt sequence for RdRp coding (ORF1b), which is conserved among AstV, as otherwise the sequence for capsid proteins (ORF2) is highly variable.1 Indeed, both regions present fundamental characteristics which impart confidence to the detection and genotyping of the lineages of AstrV, as well gaining access to the diversity of these viruses. Classic HAstV5 has not been circulating in Brazil since 1999 and here this genotype was detected in children with AGE from demarked indigenous areas which are very isolated.6 Gabay et al.14 reported in 2006 a large outbreak of AGE affecting more than 100 children under six years of age that occurred in Indigenous people from the Maxakali reserve in the State of Minas Gerais, Brazil. HAstV-2 was the sole enteropathogen detected in 26 (56%) of 46 samples analyzed. Using in vitro methods and animal models to characterize virus-host interactions, researchers have discovered several important properties of AstV, including the ability of the AstV capsid to act as an enterotoxin, disrupting the gut epithelial barrier.15 In the last few years, a lot of information has been obtained in studies involving sugars exposed in the gastrointestinal tract membrane, interacting with different etiological agents of AGE. Genetic factors and the immunological status of the host affect the expression of these sugars.15 The frequency of HAstV detected in this study could be higher because co-infection with other virus causing AGE was not considered. Therefore greater attention should be given to the young, vulnerable population living in indigenous reserves. Boa Vista is a gateway for tourists visiting the Amazon rainforest and their high circulation could bring new AstrV species. In addition, both non-classic AstVMLB2 were detected in children living in Boa Vista, RR state capital. Although conventional RT-PCR method does not have high sensitivity, it allowed showing the diversity of HAstV that circulates in the Amazon region, reinforcing the need of surveillance.
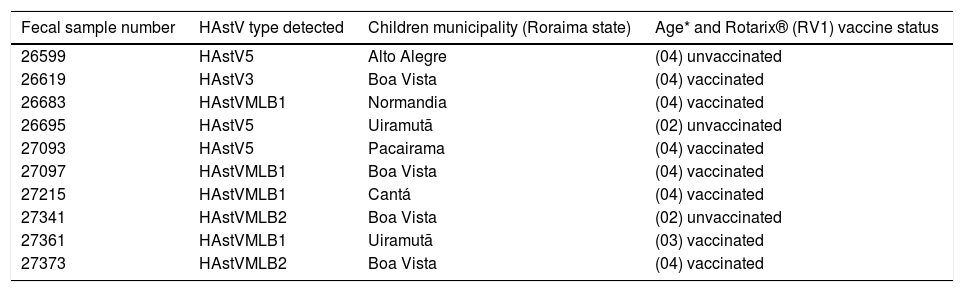
Summary of human astrovirus (HAstV) genotypes infecting children from Amazon region detected by reverse transcription polymerase chain reaction (RT-PCR). Children from the municipalities live in demarcated indigenous regions, excluding Boa Vista (capital of Roraima state).
| Fecal sample number | HAstV type detected | Children municipality (Roraima state) | Age* and Rotarix® (RV1) vaccine status |
|---|---|---|---|
| 26599 | HAstV5 | Alto Alegre | (04) unvaccinated |
| 26619 | HAstV3 | Boa Vista | (04) vaccinated |
| 26683 | HAstVMLB1 | Normandia | (04) vaccinated |
| 26695 | HAstV5 | Uiramutã | (02) unvaccinated |
| 27093 | HAstV5 | Pacairama | (04) vaccinated |
| 27097 | HAstVMLB1 | Boa Vista | (04) vaccinated |
| 27215 | HAstVMLB1 | Cantá | (04) vaccinated |
| 27341 | HAstVMLB2 | Boa Vista | (02) unvaccinated |
| 27361 | HAstVMLB1 | Uiramutã | (03) vaccinated |
| 27373 | HAstVMLB2 | Boa Vista | (04) vaccinated |
Subtitle:* The age each child, being: 1= ≤3 months old; 2= >3 months ≤6 months old; 3 = >6 months ≤1 year old; 4= >1 year ≤ 2 years old; 5= >2 years ≤5 years old. No HAstV positivity was detected among children aged 1 and 5.
A. Multiple sequence alignment of the amino acids based on partial amino acid sequences (136 aa) from the partial RNA-dependent RNA polymerase (RdRp) of human astrovirus. The AB823731.1, KJ807479 and KJ807514.1 were accessed from the National Center Biotechnology Information GenBank and used as reference samples. The GenBank access number for the sample genotyped in this study HAstV23373 is MT036255. The similarity between HAstVMLB2 samples 27373 and 27341 was >99.9%, with only one amino acid (aa) difference in the RdRp in sample number 27373 (T334A) absent in sample 27341, considering the reference sequence access number KJ807479.1. B. Phylogenetic analysis of astroviruses based on partial amino acid sequences (121 aa) of the partial RNA-dependent RNA polymerase (RdRp) from children from Boa Vista, Roraima, Brazil. Sequences were analyzed using the maximum-likelihood method and bootstrap values >70% are shown at the nodes of the tree as percentages based on 2000 replicates. The strains reported in this study are indicated by filled black circles and the reference strains of astrovirus are shown with their respective GenBank access numbers (legend: HAstV = human astrovirus; BoAstV = bovine astrovirus).
We are grateful for support from the Coordination for the Improvement of Higher Education Personnel (CAPES) “Programa de Professor Visitante no Exterior-Edital no.01/2019-Processo 88881.337140/2019-01”; The National Council for Scientific and Technological Development–CNPq number 424376/2016-4; Foundation for Research Support of the State of Rio de Janeiro-FAPERJ “Edital Carlos Chagas Filho” and Oswaldo Cruz Institute-IOC (PAEF).
Conflict of interestsAll authors declare that they have no conflict interests.
The authors acknowledge all children and their parents for making this study possible. Thanks to Professor Dr. José Francisco Luitgards Moura from UFRR, for all his support throughout the Project and for his teachings on indigenous culture. To Bruno Baroni de Moraes e Souza for the English revision and we would like to thank the staff members of “The Fiocruz Institucional Platform for DNA Sequencing (PDTIS)”.