The herein presented assay provided a bacteriological and molecular characterization of 100 samples of L. monocytogenes isolated from human (43) and food (57) sources, from several regions of Brazil, and collected between 1975 and 2013. Antigenic characterization defined 49% of serotype 4b samples, followed by 28% of serotype 1/2b, 14% of serotype 1/2c, 8% of serotype 1/2a, and 1% of serotype 3b. Both type of samples from human and food origin express the same serotype distribution. Multiplex PCR analysis showed 13 strains of type 4b with the amplification profile 4b-VI (Variant I). Virulence genes hly, inlA, inlB, inlC, inlJ, actA, plcA, and prfA were detected in all samples, highlighting a deletion of 105pb on the actA gene in 23% of serotype 4b samples. Macrorestriction profile with ApaI at PFGE showed 55 pulsotypes, with the occurrence of the same pulsotype in hospitalized patients in São Paulo in 1992 and 1997, and two other highly related pulsotypes in patients hospitalized in Rio de Janeiro in 2008. Recognized pulsotypes in listeriosis cases have also been detected in food. Thus, the prevalence of a serotype and the persistence of certain pulsotypes herald future problems.
Listeria monocytogenes is a Gram-positive bacterium, pathogenic to man and animals, usually propagated through contaminated food and frequently detected in nature. It is responsible for the clinical presentation of listeriosis and affects primarily pregnant women, newborns, elderly patients, and other patients, immunocompromised or not. It has a high mortality rate, ranging from 20% to 30%. The main clinical manifestations include gastroenteritis, meningitis, and septicemia.1,2
According to the monitoring of food-borne diseases by FoodNet in 2013, among bacterial infections, infections by L. monocytogenes had the lowest incidence (0.26/100,000), but were responsible for higher hospitalization rates (90%) and deaths (20%) in the US.3 In United States and European countries listeriosis is considered a disease of relevance to public health, while in Brazil there are limited data about its occurrence, and it is not even included in the Ministry of Health's list of mandatory notification diseases.2,4
Main determinants of virulence studied in the species are the genes hly, inlA, inlB, inlC, inlJ, actA, plcA, and prfA, involved in host cell–cell dissemination process. Loss or modification of these factors may result in attenuation of virulence of the species.5
L. monocytogenes can trigger severe clinical conditions predominantly in immunocompromised individuals, whose numbers represent a significant fraction of the world population, including Brazil. Considering this particularity, and scarce literature available, this study aims to detect the virulence profile and determine genetic relationships among samples of L. monocytogenes of human and food origins isolated in different periods.
Material and methodsBacterial strainsWe studied 100 samples of L. monocytogenes, of which 57 were of clinical origin, isolated from CSF, blood, lymph nodes, peritoneal fluid, and placenta, in the period of 1975–2013, and 43 samples from various food such as dairy products, sausages/cured meats, salads, and other ready-to-eat dishes in the period of 2001–2013. Most samples were received for confirmatory testing of the species, and came from public and private institutions from various regions of Brazil. The samples are part of the Coleção de Culturas de Listeria (CLIST, Listeria Culture Collection) at the Laboratório de Zoonoses Bacterianas (Labzoo/IOC-FIOCRUZ, Bacterial Zoonosis Laboratory), maintained by cryopreservation (in a freezer at −80°C) in Brain Heart Infusion (BHI) broth with addition of 20% glycerol.
Antigenic and biochemistry characterizationBiochemistry characterization was performed according to the Rocourt scheme.6 The characterization of serogroups and serovars was conducted according to the recommendations of Seeliger & Höhne7 using somatic and flagellar antisera produced by Labzoo/FIOCRUZ.
PCR assaysIn confirmation of the genus and species, respectively, the authors used primers for the 23S subunit gene of the ribosomal RNA and for the hly gene, which encodes listeriolysin O.8 Serotypes were confirmed by multiplex PCR assay.9 The primers and techniques mentioned in Table 1 references were used in the search for markers (hly, prfA, plcA, actA, inlA, inlB, inlC and inlJ). Extracted DNA was obtained using an extraction kit (DNeasy Blood & Tissue Kit; Qiagen, Hilden, Germany), according to the manufacturer's instructions. The amplicons generated by PCR were resolved by electrophoresis in agarose gel at 1%, with 0.5× TBE loading buffer (Bio Rad), with the exception of multiplex PCR, which used agarose gel at 2%. Gels were stained with ethidium bromide and photographed under ultra-violet light in a transilluminator (Bio Rad UV-gentm). Reference strain CDC F4555 (4b) was the positive control in all the above-mentioned PCR assays, and strains ATCC 19111 (1/2a), CDC F4976 (1/2b) and ATCC 19112 (1/2c) were used as specificity and reproducibility controls in the multiplex PCR assay.
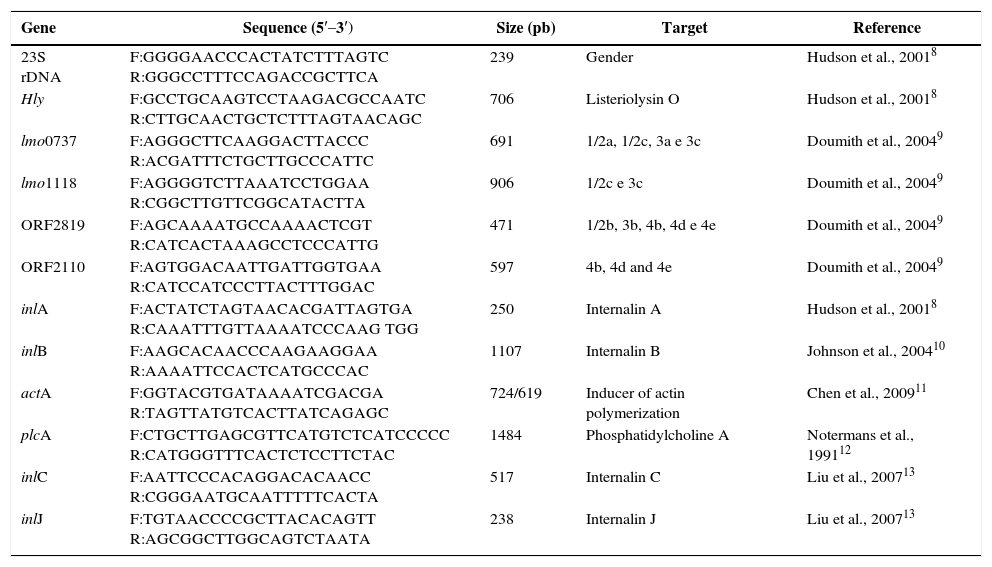
Nucleotide sequence of primers for confirmation of genus and serotype.
| Gene | Sequence (5′–3′) | Size (pb) | Target | Reference |
|---|---|---|---|---|
| 23S rDNA | F:GGGGAACCCACTATCTTTAGTC R:GGGCCTTTCCAGACCGCTTCA | 239 | Gender | Hudson et al., 20018 |
| Hly | F:GCCTGCAAGTCCTAAGACGCCAATC R:CTTGCAACTGCTCTTTAGTAACAGC | 706 | Listeriolysin O | Hudson et al., 20018 |
| lmo0737 | F:AGGGCTTCAAGGACTTACCC R:ACGATTTCTGCTTGCCCATTC | 691 | 1/2a, 1/2c, 3a e 3c | Doumith et al., 20049 |
| lmo1118 | F:AGGGGTCTTAAATCCTGGAA R:CGGCTTGTTCGGCATACTTA | 906 | 1/2c e 3c | Doumith et al., 20049 |
| ORF2819 | F:AGCAAAATGCCAAAACTCGT R:CATCACTAAAGCCTCCCATTG | 471 | 1/2b, 3b, 4b, 4d e 4e | Doumith et al., 20049 |
| ORF2110 | F:AGTGGACAATTGATTGGTGAA R:CATCCATCCCTTACTTTGGAC | 597 | 4b, 4d and 4e | Doumith et al., 20049 |
| inlA | F:ACTATCTAGTAACACGATTAGTGA R:CAAATTTGTTAAAATCCCAAG TGG | 250 | Internalin A | Hudson et al., 20018 |
| inlB | F:AAGCACAACCCAAGAAGGAA R:AAAATTCCACTCATGCCCAC | 1107 | Internalin B | Johnson et al., 200410 |
| actA | F:GGTACGTGATAAAATCGACGA R:TAGTTATGTCACTTATCAGAGC | 724/619 | Inducer of actin polymerization | Chen et al., 200911 |
| plcA | F:CTGCTTGAGCGTTCATGTCTCATCCCCC R:CATGGGTTTCACTCTCCTTCTAC | 1484 | Phosphatidylcholine A | Notermans et al., 199112 |
| inlC | F:AATTCCCACAGGACACAACC R:CGGGAATGCAATTTTTCACTA | 517 | Internalin C | Liu et al., 200713 |
| inlJ | F:TGTAACCCCGCTTACACAGTT R:AGCGGCTTGGCAGTCTAATA | 238 | Internalin J | Liu et al., 200713 |
PFGE was performed according to the PulseNetTM (Standardized Laboratory Protocol for Molecular Subtyping- http://www.cdc.gov/pulsenet/) protocol for L. monocytogenes, using the restriction enzyme ApaI (New England, Biolabs, Beverly, MA). Salmonella Braenderup H9812 (provided by the Laboratório de Enterobactérias IOC/FIOCRUZ/Brazil) was digested using the XbaI enzyme and used as a molecular weight standard. The analysis of the generated fragments was performed by BioNumerics 4.0 software (Applied Maths, Sant’Martens-Latem, Belgium). The construction of the dendrogram was prepared according to the Unweighted Pair-group Method with Averages (UPGMA) applying the Dice coefficient, with 1.5% tolerance index. For the data analysis, two PFGE profiles were considered indistinguishable when they had 100% similarity, and profiles with a difference in one single band were defined as different profiles. This criterion was adopted due to a lack of epidemiological data and the wide geographical and temporal distribution of the samples in the study. This recommendation was based on analysis by Barret et al.,14 from the network of CDC Foodborne Disease Active Surveillance Network.
Statistical analysisCategorical variables were compared using Chi-square or Fisher's exact test, as appropriate, with the aid of STATA 13.0. Results with p-value ≤0.05 were considered statistically significant.
Ethical considerationsThis work was submitted to and approved by the Research Ethics Committee of Hospital Universitário Clementino Fraga Filho, UFRJ.
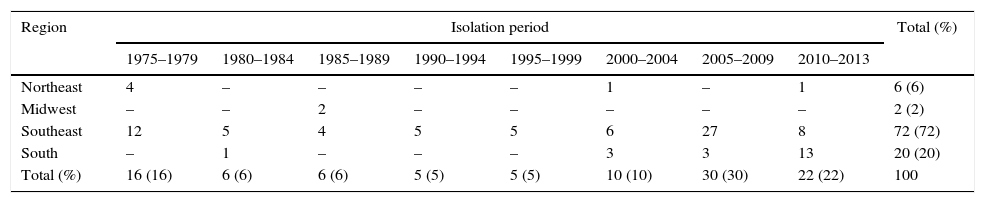
ResultsIn respect to geographical origins of the isolates, there was a predominance of samples from the States of the southeast region of the country, represented by 72 strains (45 human specimens and 27 food specimens) and the southern region (4 clinical samples and 16 food samples) followed by 6 and 2 human isolates from the Northeast and Midwest, respectively (Table 2).
Absolute and relative distribution of geographical regions of the samples and their respective periods of isolation.
| Region | Isolation period | Total (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 1975–1979 | 1980–1984 | 1985–1989 | 1990–1994 | 1995–1999 | 2000–2004 | 2005–2009 | 2010–2013 | ||
| Northeast | 4 | – | – | – | – | 1 | – | 1 | 6 (6) |
| Midwest | – | – | 2 | – | – | – | – | – | 2 (2) |
| Southeast | 12 | 5 | 4 | 5 | 5 | 6 | 27 | 8 | 72 (72) |
| South | – | 1 | – | – | – | 3 | 3 | 13 | 20 (20) |
| Total (%) | 16 (16) | 6 (6) | 6 (6) | 5 (5) | 5 (5) | 10 (10) | 30 (30) | 22 (22) | 100 |
Clinical samples were concentrated in the states of São Paulo (29 samples, 51%) and Rio de Janeiro (14, 25%), with the main sources of isolation being CSF (29, 51%) and blood (22, 39%). As for the food samples, they were mostly isolated from São Paulo (23, 54%) and Rio Grande do Sul (12, 28%), and the main source of isolation of L. monocytogenes were dairy products (20, 46%).
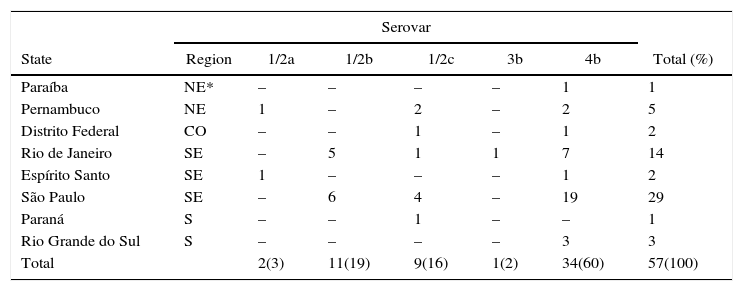
In the conventional and molecular identification of serotypes it is noteworthy to highlight the prevalence of types 4b (49%) and 1/2b (28%) in both sources, with a higher frequency of serotype 4b in clinical samples in comparison with serotype 1/2b, but without statistical significance (p>0.05) (Tables 3 and 4).
Distribution of L. monocytogenes serotypes isolated from clinical specimens according to geographical regions and states of Brazil.
| Serovar | |||||||
|---|---|---|---|---|---|---|---|
| State | Region | 1/2a | 1/2b | 1/2c | 3b | 4b | Total (%) |
| Paraíba | NE* | – | – | – | – | 1 | 1 |
| Pernambuco | NE | 1 | – | 2 | – | 2 | 5 |
| Distrito Federal | CO | – | – | 1 | – | 1 | 2 |
| Rio de Janeiro | SE | – | 5 | 1 | 1 | 7 | 14 |
| Espírito Santo | SE | 1 | – | – | – | 1 | 2 |
| São Paulo | SE | – | 6 | 4 | – | 19 | 29 |
| Paraná | S | – | – | 1 | – | – | 1 |
| Rio Grande do Sul | S | – | – | – | – | 3 | 3 |
| Total | 2(3) | 11(19) | 9(16) | 1(2) | 34(60) | 57(100) | |
NE*, Northeast; CO, Midwest; SE, Southeast; S, South.
Distribution of L. monocytogenes serotypes isolated from different food specimens according to geographical regions and states of Brazil.
| Serovar | ||||||
|---|---|---|---|---|---|---|
| State | Region | 1/2a | 1/2b | 1/2c | 4b | Total (%) |
| Rio de Janeiro | SE* | 1 | 1 | 1 | 1 | 4 |
| São Paulo | SE | 2 | 6 | 2 | 13 | 23 |
| Santa Catarina | S | – | 1 | – | 3 | 4 |
| Rio Grande do Sul | S | 2 | 3 | 3 | 4 | 12 |
| Total | 5(12) | 11(25) | 6(14) | 21(49) | 43(100) | |
SE*, Southeast; S, South.
In the 13 serotype 4b samples (11 isolates from human clinical specimens and 2 from food specimens), multiplex PCR showed the amplification profile 4b-VI (Variant I), defined by amplification of the lmo0737 band.
All genes related to virulence were detected, although the actA gene presented fragments of varying sizes. For example, there were 77 isolates (77%) with the 725pb fragment and 23 (23%) with the 619pb fragment. Interestingly, only serotype 4b isolates showed the smaller fragment of the actA gene, while serotypes 1/2a, 1/2b, and 1/2c presented the 724pb fragment.
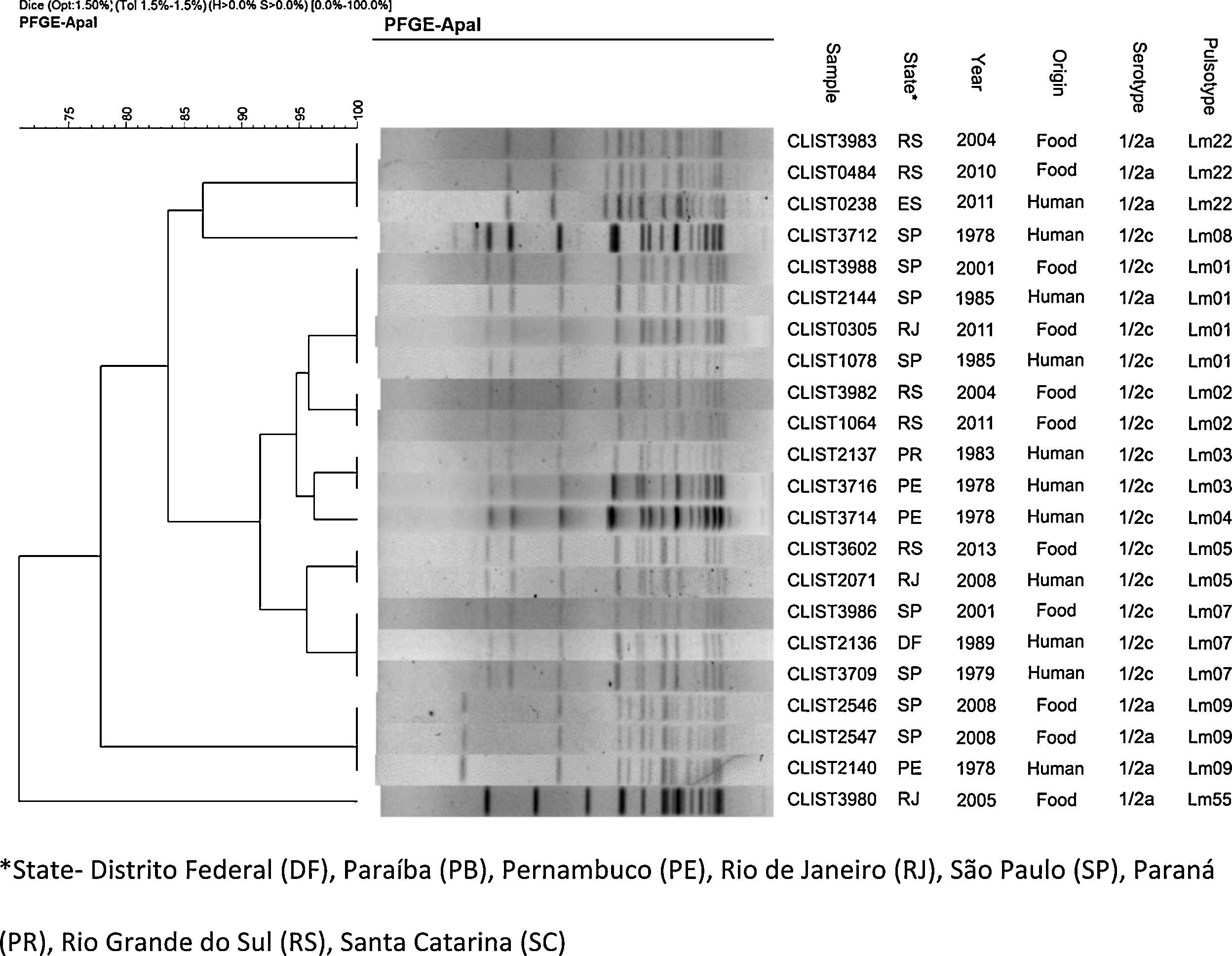
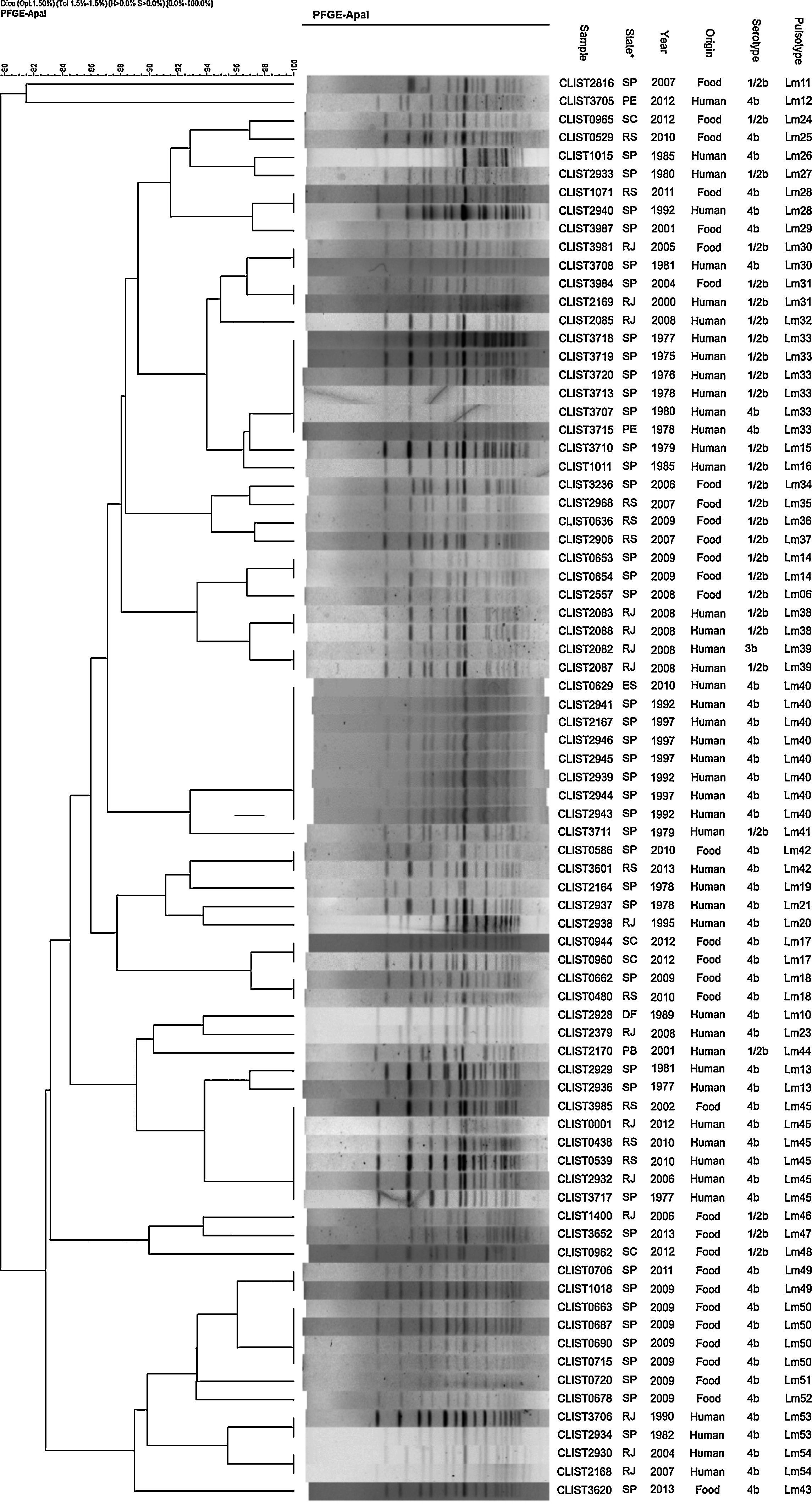
DNA profiles contained in the dendrograms (Figs. 1 and 2) divided the samples into two groups with similarity above 70%. The first group consists of serotypes 4b, 1/2b and 3b, and the second group of serotypes 1/2a and 1/2c. In total, there were 55 pulsotypes, namely: 4 pulsotypes in serotype 1/2a, 7 in serotype 1/2c, 22 in serotype 1/2b, 1 in serotype 3b, and 25 in serotype 4b. Some isolates of various serotypes are included in a same pulsotype (Lm01, Lm30, Lm33 and Lm39).
Dendrogram containing the distribution of serotypes 1/2a and 1/2c using the PFGE technique and analyzed using the Dice coefficient with 1.5% tolerance, using UPGMA. *State – Distrito Federal (DF), Paraíba (PB), Pernambuco (PE), Rio de Janeiro (RJ), São Paulo (SP), Paraná (PR), Rio Grande do Sul (RS), Santa Catarina (SC).
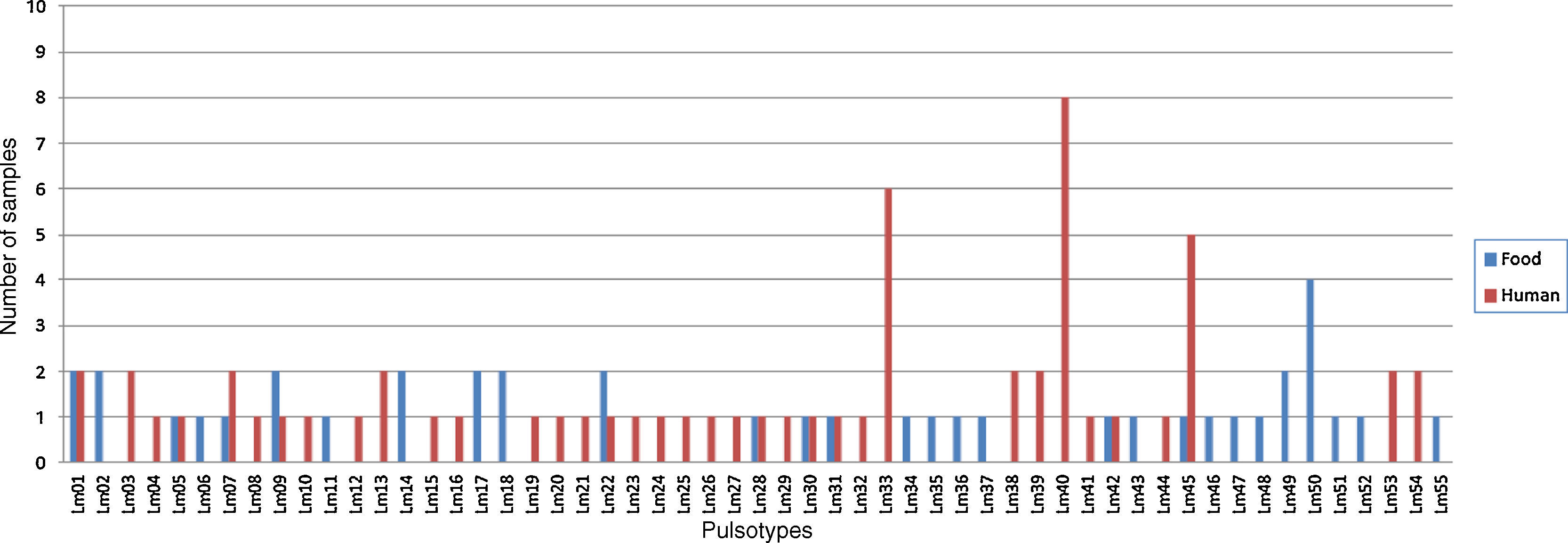
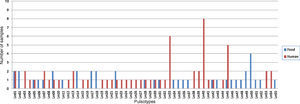
In Fig. 3, L. monocytogenes pulsotypes Lm01, Lm05, Lm07, Lm09, Lm22, Lm28, Lm30, Lm31, and Lm42 were found in samples from both human and food sources, isolated at different time points and locations. Pulsetype Lm40 (8 samples) was most often isolated, followed by pulsotypes Lm33 and Lm45 (6 samples each), Lm01 and Lm50 (4 samples each). Pulsotypes Lm49, Lm50, Lm51 and Lm52 were isolated from eight L. monocytogenes samples in salads and ready-to-eat dishes, in 2009 and 2011, in São Paulo.
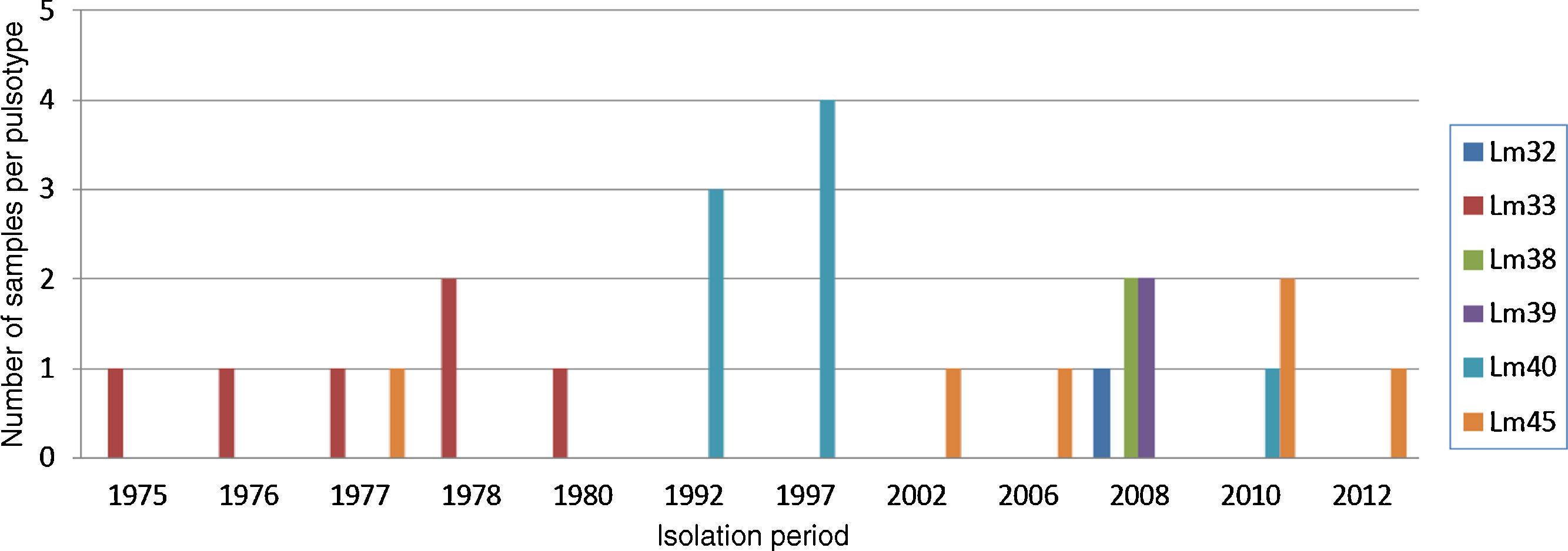
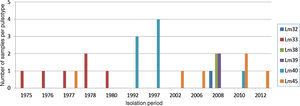
Fig. 4 shows five samples of pulsotypes Lm32, Lm39 and Lm38 isolated from human samples from a hospital in Rio de Janeiro in 2008. The Lm33 pulsotype was detected in six samples of human origin, isolated in 1975, 1976, 1977, 1978, and 1980 in São Paulo, and in 1978 in Pernambuco. Pulsotype Lm40 was present in seven samples from the state of São Paulo (1992 and 1997) and in one sample from Espírito Santo in 2010. Pulsotype Lm45 was found in five samples from cases of listeriosis in different years in the states Rio de Janeiro, Rio Grande do Sul, and São Paulo, and was also found in a dairy sample from Rio Grande do Sul.
DiscussionAntigenic and molecular identification of L. monocytogenes demonstrated a predominance of serotypes 4b and 1/2b (77%) compared to other serotypes (23%), both in clinical and food samples, consistent with numerous reports from various parts of the world, including Brazil.15–19
However, other investigations found different results, particularly in food.20,21
As such, the importance of the molecular assay developed by Doumith et al. (2004)9 is highlighted, and it is recommended as an alternative to conventional serology, which has high cost-effectiveness due to the limited availability of somatic and flagellar antisera and the need for technical expertise.21,22 Moreover, the adoption of multiplex PCR led to the recognition of an amplification profile named 4b Variant I (4b-VI) through the product lmo0737, which had only been reported in serotypes 1/2a, 3a, 1/2c and 3c.22 Leclercq et al.23 concluded that the samples with this profile belonged to at least two ST “sequence types” unrelated to MLST, and that the 4b-V1 profile does not correspond to a recent clonal emergence, suggesting that the acquisition of fragment lmo0737 would have been the result of horizontal transfer.
An important aspect to study in the pathogenesis of L. monocytogenes is the absolute presence of the analyzed virulence genes, regardless of the origin of isolates, and this is consistent with several literature citations.17 The behaviour observed in the actA gene is highlighted, wherein the samples present two distinct fragment sizes, 724pb and 619pb. As in the study by Chen et al.,11 which concentrated on isolates from food, we also detected a 105pb deletion of the actA in 23% of the serotype 4b samples. Also based on the observations by Chen et al.,11 those same isolates retained the capacity to spread to adjacent cells in cell adhesion assays and, as such, one could hypothesize the existence of other dissemination mechanisms beyond protein ActA.
On the other hand, Barbosa et al.24 suggested that such a deletion is a characteristic of the Epidemic Clone I circulating in Brazil.
In data from PFGE, some samples of different serotypes were pooled in the same pulsotype, consistent with other observations.25–27 This situation is supported in research into the evolutionary analysis of lineage I as proposed by Ragon et al.,28 indicating that serotype 4b originated from serotype 1/2b, while lineage II serotype 1/2c has serotype 1/2a ancestry. Thus, it is justifiable that different serovars can be grouped in the same pulsotype.
As a reference point in analyzing the macrorestriction profile, the characterization of nine pulsotypes consisting of strains is highlighted. They were isolated from human cases and from food sources, one to three decades before, and originating from different states (Fig. 3). In spite of such heterogeneity, the formation of recurring pulsotypes (100% similarity) was observed, suggesting their ability to adapt and circulate among possible sources of infection and, therefore, among the means of transmission. These results, addressing the same or similar aspects to those mentioned above, have been observed in other countries.19,29,30
Although we observed the presence of several pulsotypes related to sporadic human cases, our study showed the occurrence of a group of samples with the same pulsotype in a hospital in São Paulo in 1992 and 1997. Most of the samples in this group had the amplification profile 4b-VI. It is worth mentioning another cluster of cases of listeriosis in a hospital in Rio de Janeiro in 200831 and pulsotypes with 97% similarity, diverging in one band that, according to the criteria by Tenover et al.32 are highly related.
ConclusionThis investigation showed persistence and circulation of certain pulsotypes related to cases of human listeriosis, as well as in food, even after long periods and from different states. The dissemination of L. monocytogenes pulsotypes may have been facilitated by the marketing of food between different regions in Brazil. Given this situation, it is essential to have continuous monitoring of food products.
In summary, towards the prevention of listeriosis, a joint action is required by surveillance agencies, the food industry, and agencies responsible for public health. This effort must include awareness of the population to the risks inherent in each food, educational campaigns for pregnant women, elderly persons, and food handlers, as well as mandatory reporting of cases of the disease.
Conflicts of interestThe authors declare no conflicts of interest.
The authors thank Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for financial support.