Understanding the epidemiology of Streptococcus pneumoniae (S. pneumoniae) isolates is important for pneumonia treatment and prevention. This research aimed to explore the epidemiological characteristics of S. pneumoniae isolated from pediatric inpatients and outpatients during the same period.
MethodsS. pneumoniae were isolated from unsterile samples of inpatients and outpatients younger than five years old between March 2013 and February 2014. The serotypes were determined using diagnostic pneumococcal antisera. The resistance of each strain to 13 antibiotics was tested using either the E-test or the disc diffusion method. The Sequence Types (STs) were analyzed via Multilocus Sequence Typing (MLST).
ResultsThe dominant serotypes obtained from inpatients were 19F (32.9 %), 19A (20.7 %), 23F (10.7 %), 6A (10.0 %), and 14 (8.6 %), while those from outpatients were 19F (13.6 %), 23F (12.9 %), 6A (10.0 %), 6B (10.0 %), and 19A (7.9 %). The coverage rates of 13-valent Pneumococcal Conjugate Vaccine (PCV) formulations were high in both groups. The nonsusceptibility to penicillin, cefuroxime, imipenem, erythromycin, and trimethoprim-sulfamethoxazole among the inpatient isolates was 7.1 %, 92.8 %, 65.7 %, 100 %, and 85.0 %, respectively, while that among the outpatient isolates was 0.7 %, 50.0 %, 38.6 %, 96.4 %, and 65.7 %, respectively. There were 45 and 81 STs detected from the pneumococci isolated from inpatients and outpatients, respectively. CC271 was common among both inpatients and outpatients (43.6 % and 14.3 %).
ConclusionsPneumococcal vaccine-related serotypes are prevalent among both inpatients and outpatients, especially among inpatients, who exhibit more severe antibiotic resistance. Therefore, universal immunization with PCV13 would decrease the hospitalization rate due to S. pneumoniae and the antibiotic resistance rate of S. pneumoniae.
Streptococcus pneumoniae (S. pneumoniae) is a major pathogen of infectious diseases worldwide. Epidemiological data on invasive S. pneumoniae isolates are important for pneumonia treatment and prevention [1]. However, it is difficult to collect an adequate number of isolates in countries where the rate of positive bacteria cultures from invasive samples is very low. In one of our previous studies, we collected 171 invasive pneumococcal isolates from 11 hospitals between 2006 and 2008 [2]. On average, only 5.2 isolates were collected in each hospital every year. Even in recent research, only 94 invasive pneumococcal isolates were collected between 2013 and 2017 [3]. The lack of available data limits the representation of S. pneumoniae in corresponding regions. The Centers for Disease Control and Prevention (CDC) and the World Health Organization (WHO) recommend that nasopharyngeal isolates be used for the surveillance of pneumococcal epidemiology since they can be obtained in larger numbers [4]. However, in Chinese hospitals, nasopharyngeal swab collection is carried out only for some research projects and not regularly. Nasopharyngeal swab collection usually stops when the project is finished. In China and other developing countries, etiologic examination is usually performed on inpatients. Therefore, bacterial culturing of sputum (i.e., laryngohypopharynx aspirates) and bronchoalveolar lavage can be constantly carried out in daily clinical work in the inpatient department, which is favorable for continuous surveillance.
In two of our previous studies, higher rates of drug resistance and increased coverage of the 7-valent Pneumococcal Conjugate Vaccine (PCV7)-related serotype were found in isolates from the hypopharyngeal aspirates of inpatients with pneumonia than in nasopharyngeal isolates from nasopharyngeal swabs of outpatients with an upper respiratory infection [5,6]. However, these two studies were not conducted during the same period. The exact epidemiological characteristics of S. pneumoniae isolated from pediatric inpatients and outpatients during the same period could not be found in previous reports.
In the present study, clinical noninvasive S. pneumoniae isolates from inpatients and nasopharyngeal carriage isolates from outpatients visiting Beijing Children's Hospital were collected between March 2013 and February 2014. The serotype distribution, PCV coverage rate, antimicrobial resistance, and Multilocus Sequence Typing (MLST) were analyzed for the two groups of isolates.
MethodsThe subjects of the current study consisted of children younger than 5-years old at Beijing Children's Hospital between March 2013 and February 2014. Inpatients were children with pneumonia who were admitted to the Infectious Diseases Department, the Respiratory Diseases Department, or the Intensive Care Unit. The outpatients were children who visited the outpatient department with a respiratory infection. We collected 140 S. pneumoniae isolates from inpatients, which were cultured from hypopharyngeal aspirates (n = 104), bronchoalveolar lavage fluid (n = 33), nasopharyngeal swabs (n = 1) or ear discharge (n = 2). Of the 693 pneumococcal strains isolated from outpatients and cultured from nasopharyngeal swabs, 140 were randomly selected using the random number table method.
All the inpatient isolates were cultured in a clinical laboratory following a procedure similar to succedent annotation [7]. The isolates were subsequently transferred to the Microbial Laboratory for further tests. For the outpatients, nasopharyngeal swabs were collected and immediately transferred to the Microbial Laboratory. The samples were inoculated within 3‒4 h onto tryptone soy agar plates with 5 % sheep blood containing 5 g/mL gentamicin. The plates were incubated at 35 °C under a 5 % CO2 atmosphere and examined after 18‒24 h. Using the random number table method, arbitrary outpatient isolates were selected monthly to correspond to the same number of inpatient isolates.
All the isolates were identified based on their typical colony morphology using Gram staining, an optochin sensitivity test (Oxoid Company, Britain), and an Omni serum assay (Statens Serum Institut, Copenhagen, Denmark). All the isolates were stored at −80 °C in freezing tubes for further study. Only one isolate from each participant was included in the present study.
The serogroups were tested via the Quellung reaction using Pneumotest kits, and the serotypes were tested with factor antisera (Statens Serum Institute, Copenhagen, Denmark). The interpretation of the serotyping depended on the presence of capsular swelling under phase-contrast microscopy with an oil immersion lens (magnification, ×100), as described in the literature [8]. The serotype coverage rates of PCV7, PCV10, PCV13 and PCV20 were estimated by calculating the percentage of isolates that expressed the serotypes included in the vaccines.
For all the isolates, the Minimum Inhibitory Concentrations (MICs) against penicillin, erythromycin, amoxicillin-clavulanic acid, cefaclor, cefuroxime, ceftriaxone, levofloxacin, linezolid, vancomycin and imipenem were determined using E-test strips (BIOMERIEUX, France), and the antimicrobial susceptibilities to chloramphenicol, tetracycline and trimethoprim-sulfamethoxazole were determined using the Kirby-Bauer disk diffusion test (OXOID Company, Britain). The Clinical and Laboratory Standards Institute (CLSI) 2015 criteria for MICs were applied to classify isolates as susceptible, intermediate, or resistant [9]. The S. pneumoniae American Type Culture Collection strain 49,619 (ATCC 49,619), which was used as a quality control strain, was included in each set of tests to ensure the accuracy of the results. Multidrug Resistant (MDR) S. pneumoniae were defined as resistant to three or more classes of antibiotics tested in this study.
The housekeeping genes aroE, gdh, gki, recP, spi, xpt, and ddl were amplified via Polymerase Chain Reaction (PCR). The sequences of seven loci were each compared with those of all known alleles at the loci and the STs in the database of the pneumococcal MLST website (https://pubmlst.org/spneumoniae) [10]. eBURST v3 software (https://eburst.mlst. net//) was used to investigate the relationships between the isolates and to assign a Clonal Complex (CC) based on the stringent group definition of six out of seven shared alleles. Among the seven MLST loci, STs that shared six identical alleles with another ST in the group were reclassified into one group as a CC.
Ethics approval and consent to participateA parent or legal guardian of each participant signed a written informed consent document before enrollment and before any study procedure was performed. This study was reviewed and approved by the Ethics Committee of Beijing Children's Hospital affiliated with Capital Medical University. Ethical problems were not encountered in this study.
Statistical analysisThe antimicrobial resistance data were collected and analyzed using WHONET 5.6 software as recommended by the WHO, and the serotype data were collected and analyzed using Excel 2017.
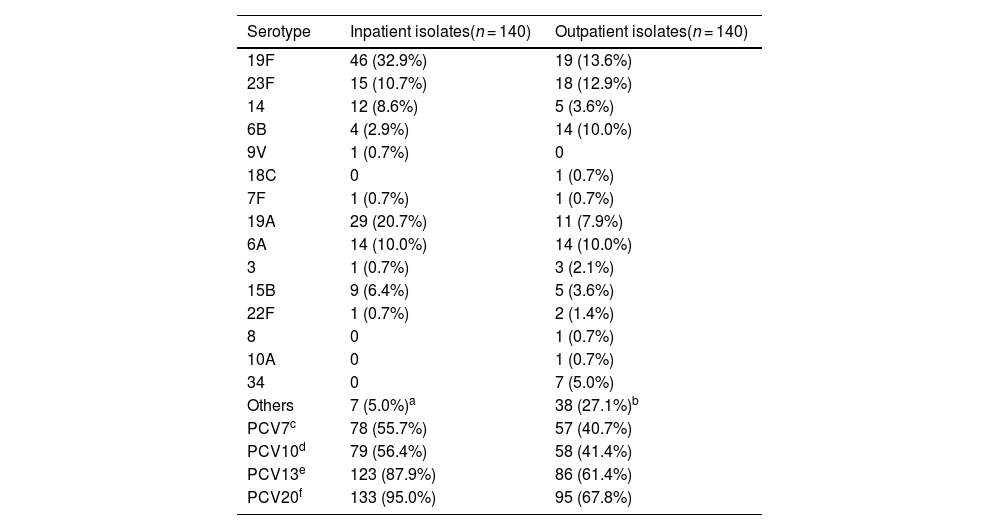
ResultsSerotype distribution and vaccine coverageThe serotype distribution and vaccine coverage of the inpatient and outpatient S. pneumoniae isolates are shown in Table 1. A serotype was included in the table when its constituent ratio was greater than 5 %. The serotype distribution was more concentrated among the inpatient isolates, in which only 15 serotypes were identified. The most common serotypes were 19F (32.9 %), 19A (20.7 %), 23F (10.7 %), 6A (10.0 %), 14 (8.6 %) and 15B (6.4 %), which accounted for 89.3 %. Moreover, 29 serotypes were identified among the outpatient isolates. The most frequent serotypes were 19F (13.6 %), 23F (12.9 %), 6A (10.0 %), 6B (10.0 %), 19A (7.9 %) and 34 (5.0 %), which accounted for 59.3 %. In addition, the serotype coverage rates of PCVs were high among inpatient isolates.
Serotype distribution and vaccine coverage rates of inpatient and outpatient S. pneumoniae isolates [n (%)].
| Serotype | Inpatient isolates(n = 140) | Outpatient isolates(n = 140) |
|---|---|---|
| 19F | 46 (32.9%) | 19 (13.6%) |
| 23F | 15 (10.7%) | 18 (12.9%) |
| 14 | 12 (8.6%) | 5 (3.6%) |
| 6B | 4 (2.9%) | 14 (10.0%) |
| 9V | 1 (0.7%) | 0 |
| 18C | 0 | 1 (0.7%) |
| 7F | 1 (0.7%) | 1 (0.7%) |
| 19A | 29 (20.7%) | 11 (7.9%) |
| 6A | 14 (10.0%) | 14 (10.0%) |
| 3 | 1 (0.7%) | 3 (2.1%) |
| 15B | 9 (6.4%) | 5 (3.6%) |
| 22F | 1 (0.7%) | 2 (1.4%) |
| 8 | 0 | 1 (0.7%) |
| 10A | 0 | 1 (0.7%) |
| 34 | 0 | 7 (5.0%) |
| Others | 7 (5.0%)a | 38 (27.1%)b |
| PCV7c | 78 (55.7%) | 57 (40.7%) |
| PCV10d | 79 (56.4%) | 58 (41.4%) |
| PCV13e | 123 (87.9%) | 86 (61.4%) |
| PCV20f | 133 (95.0%) | 95 (67.8%) |
The antimicrobial susceptibilities of the inpatient and outpatient isolates are shown in Table 2. The rate of nonsusceptibility to penicillin was high among inpatients, and the penicillin MIC50 and MIC90 values were also high. Among the inpatient isolates, the rates of nonsusceptibility to amoxicillin-clavulanic acid, imipenem, cefuroxime, cefaclor and trimethoprim-sulfamethoxazole was 7.1 %, 65.7 %, 92.8 %, 93.6 % and 85 %, respectively. The corresponding data in outpatient isolates were 0.7 %, 38.6 %, 50 %, 53.5 % and 65.7 %, respectively. All of the isolates showed high nonsusceptibility to erythromycin and tetracycline. The rate of resistance to chloramphenicol was low for both inpatient isolates and outpatient isolates. All pneumococcal isolates were susceptible to vancomycin, linezolid and levofloxacin.
Susceptibility and MICs of inpatient and outpatient S. pneumoniae isolates to 13 antibiotics.
| Antimicrobiala | Inpatient isolates (n = 140) | Outpatient solates (n = 140) |
|---|---|---|
| Penicillin | ||
| Parenteral nonmeningitis | ||
| R% | 2.1 | 0.7 |
| I% | 5 | 0 |
| Oral nonmeningitis | ||
| R% | 40.0 | 20.7 |
| I% | 54.3 | 38.6 |
| MIC range (mg/L) | 0.016‒12 | 0.004‒6 |
| MIC50 (mg/L) | 1 | 0.5 |
| MIC90 (mg/L) | 2 | 1.5 |
| Amoxicillin-clavulanic acid | ||
| R% | 0.7 | 0 |
| I% | 6.4 | 0.7 |
| MIC range (mg/L) | 0.016‒12 | 0.016‒3 |
| MIC50 (mg/L) | 1 | 0.38 |
| MIC90 (mg/L) | 2 | 1.5 |
| Cefuroxime | ||
| R% | 87.1 | 43.6 |
| I% | 5.7 | 6.4 |
| MIC range (mg/L) | 0.023‒48 | 0.016‒12 |
| MIC50 (mg/L) | 3 | 0.5 |
| MIC90 (mg/L) | 8 | 4 |
| Cefaclor | ||
| R% | 90.7 | 51.4 |
| I% | 2.9 | 2.1 |
| MIC range (mg/L) | 0.19->256 | 0.125->256 |
| MIC50 (mg/L) | 24 | 3 |
| MIC90 (mg/L) | 64 | 96 |
| Ceftriaxone | ||
| R% | 1.4 | 0.7 |
| I% | 17.9 | 10.7 |
| MIC range (mg/L) | 0.012‒4 | 0.006‒3 |
| MIC50 (mg/L) | 0.75 | 0.38 |
| MIC90 (mg/L) | 1.5 | 1.5 |
| Imipenem | ||
| R% | 0.7 | 0 |
| I% | 65 | 38.6 |
| MIC range (mg/L) | 0.008‒2 | 0.004‒0.38 |
| MIC50 (mg/L) | 0.19 | 0.094 |
| MIC90 (mg/L) | 0.25 | 0.25 |
| Vancomycin | ||
| MIC range (mg/L) | 0.19‒0.5 | 0.25‒1 |
| MIC50 (mg/L) | 0.5 | 0.5 |
| MIC90 (mg/L) | 0.5 | 0.5 |
| Linezolid | ||
| MIC range (mg/L) | 0.38‒2 | 0.25‒2 |
| MIC50 (mg/L) | 0.75 | 1 |
| MIC90 (mg/L) | 1 | 1.5 |
| Levofloxacin | ||
| MIC range (mg/L) | 0.38‒2 | 0.38‒2 |
| MIC50 (mg/L) | 0.75 | 0.75 |
| MIC90 (mg/L) | 1 | 1 |
| Erythromycin | ||
| R% | 100 | 96.4 |
| MIC range (mg/L) | 4‒>256 | 0.094‒256 |
| MIC50 (mg/L) | >256 | >256 |
| MIC90 (mg/L) | >256 | >256 |
| Tetracycline | ||
| R% | 92.9 | 93.6 |
| I% | 4.2 | 1.4 |
| Chloramphenicol | ||
| R% | 4.3 | 10.7 |
| Trimethoprim-sulfamethoxazole | ||
| R% | 85 | 65.7 |
| I% | 2.1 | 11.4 |
Breakpoints were based on the 2015 criteria of the CLSI. The breakpoint of penicillin was based on the parenteral nonmeningitis (intermediate [4 mg/L], resistant [≥ 8 mg/L]), and oral nonmeningitis (intermediate [0.12‒1 mg/L], resistant [≥ 2 mg/L]). The breakpoint of ceftriaxone was based on only parenteral nonmeningitis (intermediate [2 mg/L], resistant [≥ 4 mg/L]). The breakpoint of amoxicillin-clavulanic acid was based on nonmeningitis (intermediate [2 mg/L], resistant [≥ 4 mg/L]). The breakpoint of cefuroxime was based on parenteral nonmeningitis (intermediate [2 mg/L], resistant [≥ 4 mg/L]). The breakpoint of cefaclor was (intermediate [2 mg/L], resistant [≥ 4 mg/L]). The breakpoint of vancomycin was defined as susceptibility ≤ 1 mg/L. The breakpoint of linezolid was defined as susceptibility ≤ 2 mg/L. The breakpoint of erythromycin was (intermediate [0.5 mg/L], resistant [≥ 1 mg/L]). The breakpoint of levofloxacin was (intermediate [14‒16 mg/L], resistant [≤ 13 mg/L]). The breakpoint of tetracycline was (intermediate [25‒27 mg/L], resistant [≤ 24 mg/L]). The breakpoint of chloramphenicol was defined as resistant [≤ 20 mg/L]. The breakpoint of trimethoprim-sulfamethoxazole was (intermediate [16‒18 mg/L], resistant [≤ 15 mg/L]).
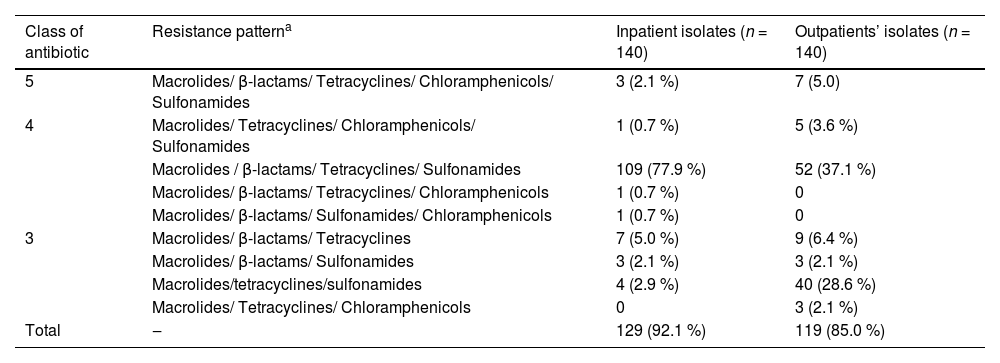
The multidrug resistance patterns of the pneumococcal isolates are shown in Table 3. Approximately 92.1 % (129/140) of the inpatient isolates and 85.0 % (119/140) of the outpatient isolates were MDR S. pneumoniae isolates. The most prevalent antibiotic resistance pattern of macrolides/β-lactams/tetracyclines/sulfonamides was observed in both inpatient and outpatient isolates.
Multidrug resistance patterns of inpatient and outpatient S. pneumoniae isolates [n (%)].
| Class of antibiotic | Resistance patterna | Inpatient isolates (n = 140) | Outpatients’ isolates (n = 140) |
|---|---|---|---|
| 5 | Macrolides/ β-lactams/ Tetracyclines/ Chloramphenicols/ Sulfonamides | 3 (2.1 %) | 7 (5.0) |
| 4 | Macrolides/ Tetracyclines/ Chloramphenicols/ Sulfonamides | 1 (0.7 %) | 5 (3.6 %) |
| Macrolides / β-lactams/ Tetracyclines/ Sulfonamides | 109 (77.9 %) | 52 (37.1 %) | |
| Macrolides/ β-lactams/ Tetracyclines/ Chloramphenicols | 1 (0.7 %) | 0 | |
| Macrolides/ β-lactams/ Sulfonamides/ Chloramphenicols | 1 (0.7 %) | 0 | |
| 3 | Macrolides/ β-lactams/ Tetracyclines | 7 (5.0 %) | 9 (6.4 %) |
| Macrolides/ β-lactams/ Sulfonamides | 3 (2.1 %) | 3 (2.1 %) | |
| Macrolides/tetracyclines/sulfonamides | 4 (2.9 %) | 40 (28.6 %) | |
| Macrolides/ Tetracyclines/ Chloramphenicols | 0 | 3 (2.1 %) | |
| Total | ‒ | 129 (92.1 %) | 119 (85.0 %) |
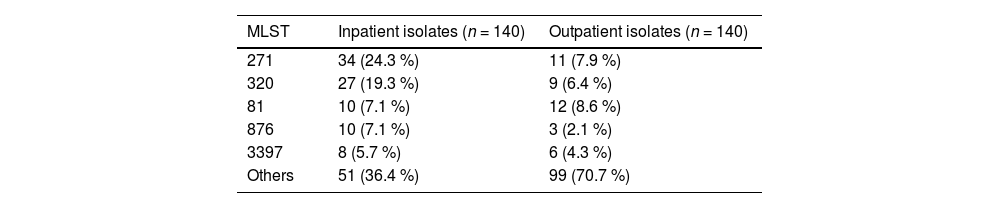
The MLSTs of the inpatient and outpatient isolates are shown in Table 4. Among the inpatient isolates, forty-five STs were detected, and the predominant STs were ST271 (24.3 %, 34/140), ST320 (19.3 %, 27/140), ST81 (7.1 %, 10/140), ST876 (7.1 %, 10/140) and ST3397 (5.7 %, 8/140), which were mainly associated with serotypes 19F, 19A, 23F, 14 and 15B, respectively. Eighty-one STs were detected among the outpatient isolates, and the predominant STs were ST81 (8.6 %, 12/140), ST271 (7.9 %, 11/140), and ST320 (6.4 %, 9/140), which were associated with serotypes 23F, 19F, and 19A, respectively.
A population snapshot of the S. pneumoniae isolates determined using eBURST analysis is shown in Fig. 1. The eBURST analysis revealed five CCs and twenty-four singletons among the inpatient isolates, and fifteen CCs and forty-two singletons among the outpatient isolates. CC271 was the most common CC among both the inpatient isolates and outpatient isolates (43.6 % and 14.3 %, respectively).
The size of the dot was proportional to the number of strains included in the ST. The line represents that there was a single site mutation between the two clones. The black STs represent the isolates from inpatients, the green STs represent the isolates from outpatients, and the purple STs represent the isolates from the two groups of patients.
DiscussionThis study described the serotype distribution, antimicrobial resistance, and molecular characteristics of S. pneumoniae isolated from inpatients and outpatients during the same period. The data showed that the serotypes 19F, 19A, 23F, and 6A were common in both inpatient and outpatient isolates, a result in line with those of previous studies regarding inpatients with pneumonia and outpatients with upper respiratory infection [6,7]. Among the inpatient isolates, only fifteen serotypes were identified, with the combined rate of serotypes 19A and 19F accounting for 53.6 %, while twenty-nine serotypes were identified among the outpatient isolates, with serotypes 19A and 19F accounting for 21.5 %. Serotype distribution was more concentrated and vaccine-related serotypes were more common among inpatient isolates. Universal immunization can potentially decrease hospitalization rate. More comprehensive serotypes were identified among the outpatient isolates, suggesting that outpatient isolates could contribute to a more comprehensive S. pneumoniae serotype distribution diagram, which could provide additional information for the evaluating serotype replacement after the PCV immunization.
Serotypes 19F and 19A were prevalent in both inpatient and outpatient isolates. These two serotypes are very common in invasive pneumococcal diseases [11–13], suggesting that they may frequently cause severe pneumococcal infections. Moreover, these two serotypes also exhibited high levels of antibiotic resistance. A previous study revealed that half of the penicillin-resistant isolates were serotype 19F, and that 47 of the serotype 19A isolates were nonsusceptible to cefuroxime, including 42 (89.4 %) that were resistant to cefuroxime [2].. A study on serotype 19F of S. pneumoniae showed that the rate of nonsusceptibility to cefaclor and cefuroxime increased from 14.2 % in 1997‒1998 to more than 80 % in 2010 [14]. One study from Bulgaria reported that the multidrug resistance of serotype 19A was 82.7 % [15]. In addition, a study revealed that serotype 19A was the major MDR serotype (38.5 %) in the USA [16]. A seminal multicenter survey on antibiotic use in five tertiary children's hospitals showed that cephalosporins were used more often [17]. These results suggest that the prevalence of serotypes 19F and 19A among inpatient isolates and outpatient isolates may be related to the selective pressure of antibiotics. Several studies have proven that current PCVs are highly effective at reducing the pneumococcal colonization rate, disease morbidity and prevalence of drug-resistant infections [18–21]. As the major serotypes in both inpatients and outpatients, serotypes 19F and 19A were included in PCV13. Universal immunization with PCV13 would decrease the antibiotic resistance of S. pneumoniae in both inpatients and outpatients.
In this study, we found that the serotype coverage rates of PCVs were high for both inpatient and outpatient isolates. The serotype coverage rates of PCV20 were greater than those of PCV13 for both inpatient isolates and outpatient isolates, especially for inpatient isolates. This finding is consistent with recent research data (93.6 % and 93.9 %, respectively) [22,23]. It is foreseeable that the immunization with PCV could reduce the prevalence of pneumococcal diseases caused by the current prevalent serotypes in China. Research from developed countries has shown that much of the remaining Invasive Pneumococcal Disease (IPD) and Pneumococcal Disease (PD) burdens are due to the additional serotypes included in PCV20. The implementation of PCV20 would further reduce the pneumococcal disease burden [24–26]. After further evaluation of its safety, immunogenicity, and effectiveness in pediatric populations, in April 2023, PCV20 received its first pediatric approval, in the USA [27]. The United Kingdom and Canada also estimated that the cost-Effectiveness of PCV20 would be greater than that of PCV13 [28,29]. PCV13 was approved in China in November 2016 and was available in the country in May 2017. Various pathogens, including bacteria, viruses, and Mycoplasma pneumoniae, etch, can cause pneumonia. For the above reason, the disease burden of PCV in preventing pneumonia is difficult to estimate, and the effectiveness achieved after vaccination is also difficult to observe significantly. Therefore, PCV13 has not been introduced into the national immunization program. In addition, because of its high price, the immunization rate of PCV13 among Chinese children is low.
Among the inpatient isolates, 94.3 % had a penicillin nonsusceptibility rate according to the oral penicillin breakpoint (intermediate [0.12‒1 mg/L], resistant [≥2 mg/L]), and the corresponding rate for the outpatient isolates was 59.3 %. Both figures were significantly greater than those for developed countries (22.4 %) [12]. A systematic literature review of the prevalence, mechanisms, and clinical implications of S. pneumoniae resistance revealed that there has been a steady decline in the susceptibility of S. pneumoniae to commonly used beta-lactams [30]. The rate of nonsusceptibility to penicillin, amoxicillin-clavulanic acid, imipenem, cefuroxime, cefaclor and trimethoprim-sulfamethoxazole, and the multidrug resistance rate were both high among the inpatient isolates. These findings suggested that antibiotic resistance was different between inpatients and outpatients, so different empirical therapies should be used under dissimilar conditions. The Chinese Medical Association recommended different treatment and antimicrobial programs for inpatients and outpatients with infectious S. pneumoniae according to the severity of their disease instead of pathogen resistance [31], and the present study provides an epidemiological reference for such an antimicrobial program. According to a seminal multicenter survey on antibiotic use in five tertiary children's hospitals, penicillins, macrolides, and cephalosporins were prescribed more often in the outpatient department, while second and third generation cephalosporins were prescribed more often in the inpatient department, with the fourth generation cephalosporins and carbapenems also being administered [17].. The present study indicates that the use of an empirical antibiotic program should be differ between the inpatients and outpatients.
Similar to the serotype distribution, STs were more concentrated among inpatient isolates. Forty-five STs were identified among the inpatient isolates, with ST271, ST320, ST81, ST876, and ST3397 being the most common STs. Eighty-one STs were detected among the outpatient isolates, with ST81, ST271, and ST320 being the most common STs. The MLST of two groups of isolates was consistent with that of a previous study [32]. All of the aforementioned results indicate that the microbiological characteristics were different between inpatients and outpatients. Twenty-five STs were identified only among the inpatient isolates. Inpatients have more serious illnesses, which means that some of the STs may lead to serious pneumococcal diseases. One study from India revealed that the STs of invasive isolates were different from those of nasopharyngeal carriage isolates. Among invasive isolates, ST63, ST4219, ST236, ST11921 and ST3135 were the most common, while among nasopharyngeal carriage isolates, ST4894, ST1701 and ST236 were the most common [33]. A survey of epidemiological data revealed the associations of the ancestral type of the capsular regulatory genome with carriage and the associations of laterally transferred sequences with invasive disease isolates [34]. STs may be related to the pathogenicity of S. pneumoniae.
CC271, the most common CC in both groups, included ST271, ST320, ST236, and ST4314. ST271 and ST320, which were more prevalent among the inpatient isolates, were associated with serotypes 19F and 19A, respectively. A systematic review of studies on invasive pneumococcal disease also revealed that CC271, which is associated with high-level antibiotic resistance, was the major CC in mainland China [35]. According to the data of the Pneumococcal Molecular Epidemiology Network (https://pubmlst.org/spneumoniae) ST320, belonges to Taiwan19F-14, which is a major antibiotic-resistant strain worldwide. This was in concordance with a study in Malaysia. The incidence of ST236, ST271 and ST320 in CC271 were reportedly associated with the MDR clone Taiwan19F-14 [36]. A Western China study revealed that CC271 isolates expressed more of mef (A/E), erm(B), lytA, rlrA and sipA than non-CC271 isolates [37]. The prevalence of resistant CC might be the main reason for the high levels of antimicrobial resistance among inpatients and outpatients.
This study has several limitations. This was a single-center study. For several reasons, clinical background information such as sex, diagnosis, and treatment was unavailable. We described only the epidemiological characteristics of S. pneumoniae isolated from inpatients and outpatients. Longitudinal and multicenter surveillance of pneumococcal isolates with additional clinical background information is necessary to confirm the present results and evaluate the influence of universal immunization with PCVs in the future.
In conclusion, pneumococcal vaccine-related serotypes with strong antibiotic resistance were still prevalent among both inpatients and outpatients. The serotypes and STs were more concentrated among the inpatient isolates. With universal immunization, the hospitalization rate could decrease. Antibiotic resistance was different between inpatients and outpatients; thus, different empirical therapies should be administered under dissimilar conditions. The present study described the serotype distribution, antimicrobial resistance, and molecular characteristics of S. pneumoniae isolated from inpatients and outpatients during the same period, which might be helpful for understanding the epidemiology of S. pneumoniae in Beijing and provide additional information for the universal immunization with PCV 13.