The rapid spread of plasmid-mediated mcr-1 gene has become a worldwide concern since it confers resistance to polymyxins considered as a last resource for treatment of infections caused by multidrug-resistant Gram-negative bacilli.1 Despite its low frequency in Brazil, the mcr-1 gene has been reported in Escherichia coli2 and Klebsiella pneumoniae3 clinical isolates. In this study, we report a case of bloodstream infection by mcr-1-harboring E. coli in southern Brazil.
The presence of mcr-1 gene was investigated in 340 polymyxin-resistant Gram-negative bacilli clinical isolates (Enterobacteriaceae, Acinetobacter baumannii and Pseudomonas aeruginosa) collected between August 2015 and January 2018 in a university hospital located in Santa Maria, Rio Grande do Sul, Brazil. Among clinical isolates, one E. coli harbored the mcr-1 gene isolated from blood cultures in September 2017. The patient was a 59-year-old woman with malignant neoplasm of middle third of esophagus and intrahepatic cholangiocarcinoma admitted to the hospital for a transthoracic esophagectomy. Four days after the procedure, the patient was transferred to the Adult Intensive Care Unit (ICU) due to hemodynamic instability and ventilatory discomfort. Empirical treatment with ceftriaxone (1 g 12/12 h) and metronidazole (1.5 mg/day) was initiated. Laboratory tests revealed leukocytosis (27,115 ± 7684.20/mm³) with left shift (8997.75 ± 1619.14/mm³ immature leukocytes) and increased C-reactive protein (>25 mg/dL; reference value: <0.3 mg/dL). E. coli isolate was recovered in two blood cultures from different peripheral sites and Klebsiella pneumoniae carbapenemase-producing K. pneumoniae (KPC-Kp) isolate in a rectal swab culture for epidemiological surveillance. Based on antimicrobial susceptibility testing (Table 1) by the VITEK®2 system (bioMérieux, Marcy-l’Étoile, France), the antibiotic regimen was switched to amikacin (250 mg 12/12 h) and meropenem (1 g 8/8 h). Nine days after transthoracic esophagectomy, the patient had complications such as septic shock, peritonitis, surgical wound dehiscence and died.
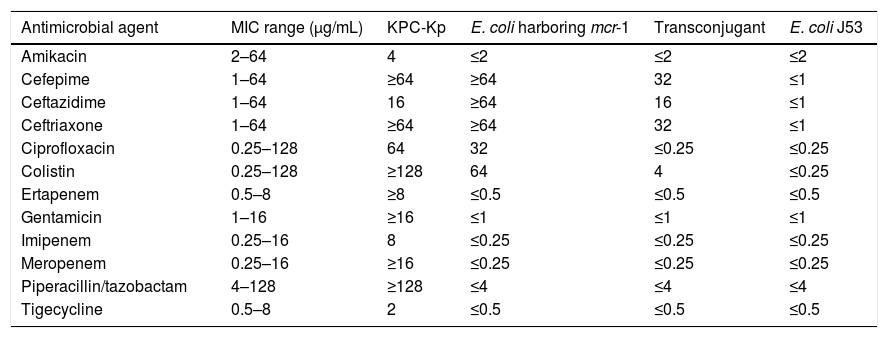
Antimicrobial susceptibility profile of KPC-Kp, E. coli harboring mcr-1, transconjugant and E. coli J53.
| Antimicrobial agent | MIC range (µg/mL) | KPC-Kp | E. coli harboring mcr-1 | Transconjugant | E. coli J53 |
|---|---|---|---|---|---|
| Amikacin | 2–64 | 4 | ≤2 | ≤2 | ≤2 |
| Cefepime | 1–64 | ≥64 | ≥64 | 32 | ≤1 |
| Ceftazidime | 1–64 | 16 | ≥64 | 16 | ≤1 |
| Ceftriaxone | 1–64 | ≥64 | ≥64 | 32 | ≤1 |
| Ciprofloxacin | 0.25–128 | 64 | 32 | ≤0.25 | ≤0.25 |
| Colistin | 0.25–128 | ≥128 | 64 | 4 | ≤0.25 |
| Ertapenem | 0.5–8 | ≥8 | ≤0.5 | ≤0.5 | ≤0.5 |
| Gentamicin | 1–16 | ≥16 | ≤1 | ≤1 | ≤1 |
| Imipenem | 0.25–16 | 8 | ≤0.25 | ≤0.25 | ≤0.25 |
| Meropenem | 0.25–16 | ≥16 | ≤0.25 | ≤0.25 | ≤0.25 |
| Piperacillin/tazobactam | 4–128 | ≥128 | ≤4 | ≤4 | ≤4 |
| Tigecycline | 0.5–8 | 2 | ≤0.5 | ≤0.5 | ≤0.5 |
Resistance to colistin was verified by broth microdilution according to EUCAST (http://www.eucast.org) and confirmed by polymyxin-NP test.4 Presence of the mcr-1 gene was verified by conventional PCR using specific primers5 and detected only in the E. coli isolate, being confirmed by Sanger sequencing. Species identification of the mcr-1-positive isolate was confirmed using MALDI-TOF MS system (Bruker Daltonics, Germany). To assess the transfer ability of the mcr-1 gene, conjugation experiment with azide-resistant E. coli J53 was performed. We were able to obtain one tranconjugant carrying mcr-1 gene and it was selected on Luria-Bertani agar supplemented with 150 µg/mL sodium azide and 2 µg/mL colistin. The transconjugant presented elevated MIC for colistin in comparison with E. coli J53.
This study reports the first detection of mcr-1 gene in Santa Maria, RS, Brazil and emphasizes the need to strengthen hospital infection prevention and control measures to prevent its spreads.
Financial supportNone reported.
Conflicts of interestThe authors declare no conflicts of interest.
The authors would like to thank Silvana Silveira Coelho and Roberta Filipini Rampelotto for technical support.