Mycobacterium tuberculosis (MTB) adopts a special survival strategy to overcome the killing mechanism(s) of host immune system. Amongst the many known factors, small heat shock protein 16.3 (sHSP16.3) of MTB encoded by gene hspX has been reported to be critical for the survival of MTB. In the present study, the effect of recombinant murine interferon-gamma (rmIFN-γ) and recombinant murine interleukin-10 (rmIL-10) on the expression of gene hspX of MTB in murine macrophage RAW264.7 has been investigated. By real-time RT-PCR, it was observed that three increasing concentrations (5, 25 and 50ng/ml) of rmIFN-γ significantly up-regulated the expression of hspX whereas similar concentrations of rmIL-10 (5, 25 and 50ng/ml) significantly down-regulated the hspX expression. This effect was not only dependent on the concentration of the stimulus but this was time-dependent as well. A contrasting pattern of hspX expression was observed against combinations of two different concentrations of rmIFN-γ and rmIL-10. The study results suggest that rIL-10 mediated down-regulation of hspX expression, in the presence of low concentration of rIFN-γ, could be used as an important strategy to decrease the dormancy of MTB in its host and thus making MTB susceptible to the standard anti-mycobacterial therapy used for treating tuberculosis. However, as these are only preliminary results in the murine cell line model, this hypothesis needs to be first validated in human cell lines and subsequently in animal models mimicking the latent infection using clinical isolates of MTB before considering the development of modified regimens for humans.
Survival of Mycobacterium tuberculosis (MTB) in human host is through battery of complex mechanisms that protect the pathogen from host's killing response.1 Amongst the several factors known to be associated with survival of MTB, small heat shock protein 16.3 (sHSP16.3) has been found indispensible for the survival of MTB.2 An in vitro study suggests that this protein acts as a molecular chaperone preventing thermal aggregation and forms a specific oligomeric structure. The chaperone activity of sHSP16.3 is temperature dependent and ATP independent.3,4
sHSP16.3 was initially identified as a 14kDa immunodominant antigen5,6 but subsequently it was found that this protein accumulates in greater quantity during stationary-phase of MTB,2 however, its turnover rate was found low.7 It is a major membrane protein.8 Functional characterization studies revealed that this protein is composed of 144 amino acid residues with apparent molecular mass of 16,277 and showed a marked homology with a member of α-crystallin family or α-heat shock protein (α-HSP) superfamily.9 It is encoded by gene hspX (acr, Rv2031c).10 In addition to playing a vital role as one of rescuers during denaturation of one of its building blocks (i.e. protein), sHSP16.3 has also been found to be vital for MTB's virulence11; cell wall thickening,12 and maintenance of long-term dormancy.2,7 In a knock out study, it was found that deletion of hspX gene speeds up the growth of MTB both in mouse and macrophage models.13
The importance of sHSP16.3 in the survival of MTB has been well established. However, no published data are available on inhibition of synthesis of this vital protein by suppressing the expression of its encoding gene hspX. Studies carried out earlier showed that the presence of oxygen,2,14 reactive nitrogen intermediates (RNIs),15 and vitamin C16 increase the expression of hspX of MTB in vitro. The merit of down-regulation of hspX expression needs to be explored in its possible role to prevent dormancy, allow multiplication of the parasite so that anti-mycobacterial agents used for therapy of tuberculosis can act effectively, thus, eliminating the organisms.
Cytokines are a diverse class of regulatory proteins or glycoproteins produced by white blood cells and a number of other cells in the body. Cytokines are involved in a broad array of biological functions. Interferon-gamma (IFN-γ) and interleukin-10 (IL-10) which are representative of Th1 and Th2 cytokines, respectively, are potent immunomodulators that play a vital role in modulation of the immune response of macrophage in opposite manner.17 IFN-γ is a major activator of macrophage. Besides, it mediates production of effector molecules such as reactive nitrogen intermediates (RNIs) via activation of tumor necrosis factor-alpha (TNF-α) cascades, resulting in killing of intracellular pathogens.18 On the other side, IL-10 is a key suppressor of macrophages and exerts its anti-inflammatory effects mainly on the phagocytic as well as antigen-presenting cells (APCs). It profoundly inhibits the production of Th1 cytokines and other co-stimulatory molecules as well as deactivates the intracellular anti-bacterial mechanisms, helping the intracellular pathogens to survive inside the host.19
The present study aimed to investigate the effect of rmIFN-γ and rmIL-10 on the expression of hspX gene of MTB grown in murine macrophage RAW264.7. After conducting the real-time RT-PCR analysis, the study provides some in vitro evidence of down-regulation of hspX of MTB by rmIL-10 in the presence of low concentration of rmIFN-γ.
Materials and methodsCulture of MTB and macrophagesH37Rv strain of MTB (ATCC, USA) was cultured and continuously maintained in Middlebrook 7H9 medium (Difco) supplemented with 10% ADC (BD), 0.2% glycerol and 0.05% Tween-80. Murine macrophage RAW264.7 (ATCC) was cultured in DMEM (Gibco) supplemented with 10% (v/v) heat inactivated FBS (Hyclone), 1.46g/l l-glutamine (Sigma), 2.3g/l HEPES (Gibco), 3.7g/l NaHCO3 (Sigma) and 10ml/l of Pen Strep (Gibco) and maintained at 37°C in a 5% CO2 humidified incubator.
MTB infection in macrophagesAn exponentially grown culture of MTB (OD600 0.6) was used to infect the 1.5×106 RAW264.7 cells at 1:15 multiplicity of infection (MOI) in 90×100mm Cell Culture Dish (Corning) containing antibiotic free complete DMEM. Single cell suspension of MTB was prepared. After eight hours of infection, cells were washed twice with warm antibiotic free DMEM and additionally treated with 0.2mg/ml amikacin sulfate (Sigma) for two hours to remove any remaining extracellular microbes. Subsequently, infected cells were replenished with fresh complete antibiotic added DMEM and incubated for the two time-points (i.e. 12h and 24h) in the presence or absence of various concentrations of rmIFN-γ (R&D Systems) and rmIL-10 (R&D Systems).
RNA isolation, cDNA synthesis and real-time RT-PCRAt specified time-points, MTB infected RAW264.7 cells were harvested and total RNA isolation was carried out using GeneJET™ RNA purification kit (Fermentas). A total of 1μg RNA was transcribed into cDNA by following the protocol of Maxima™ first strand cDNA synthesis kit (Fermentas). The isolated RNA was treated with RNase-free DNase I (Fermentas). Real-time RT-PCR analysis for the detection of hspX expression was performed using premixed SYBR Green I reaction mixture and following primers pairs (hspX: F: 5′-CGACAAGGACGTCGACATTA-3′, R: 5′-CCTTGTCGTAGGTGGCCTTA-3′; 16S rRNA: F:5′-GCGCAGATATCAGGAGGAAC-3′, R: 5′-AAGGAAGGAAACCCACACCT-3′) on the CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad). The amplification procedure involved initial denaturation at 94°C for 5min followed by 35 cycles of denaturation at 94°C for 45s, annealing of primers at 65°C for 30s and primer extension at 72°C for 45s. After completion of the 35° cycle, the extension reaction was continued for another 5min at 72°C. 16S rRNA was used an internal control for the analysis.
Statistical analysisOne-way ANOVA (repeated measures) and Bonferroni's post hoc tests were performed to determine the statistical significance among various groups using GraphPad Prism® 5.0 software. p-Values were considered significant if p<0.05.
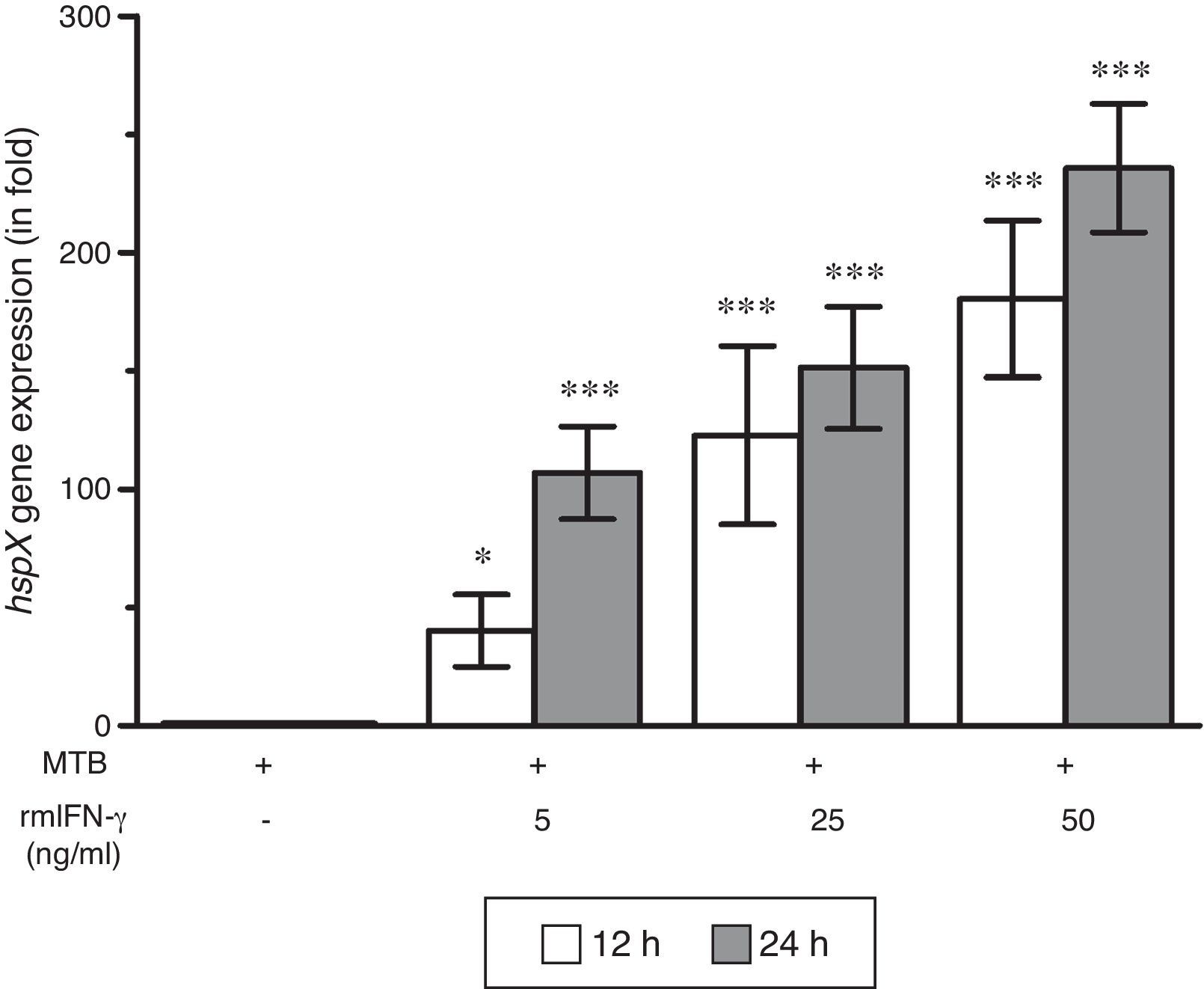
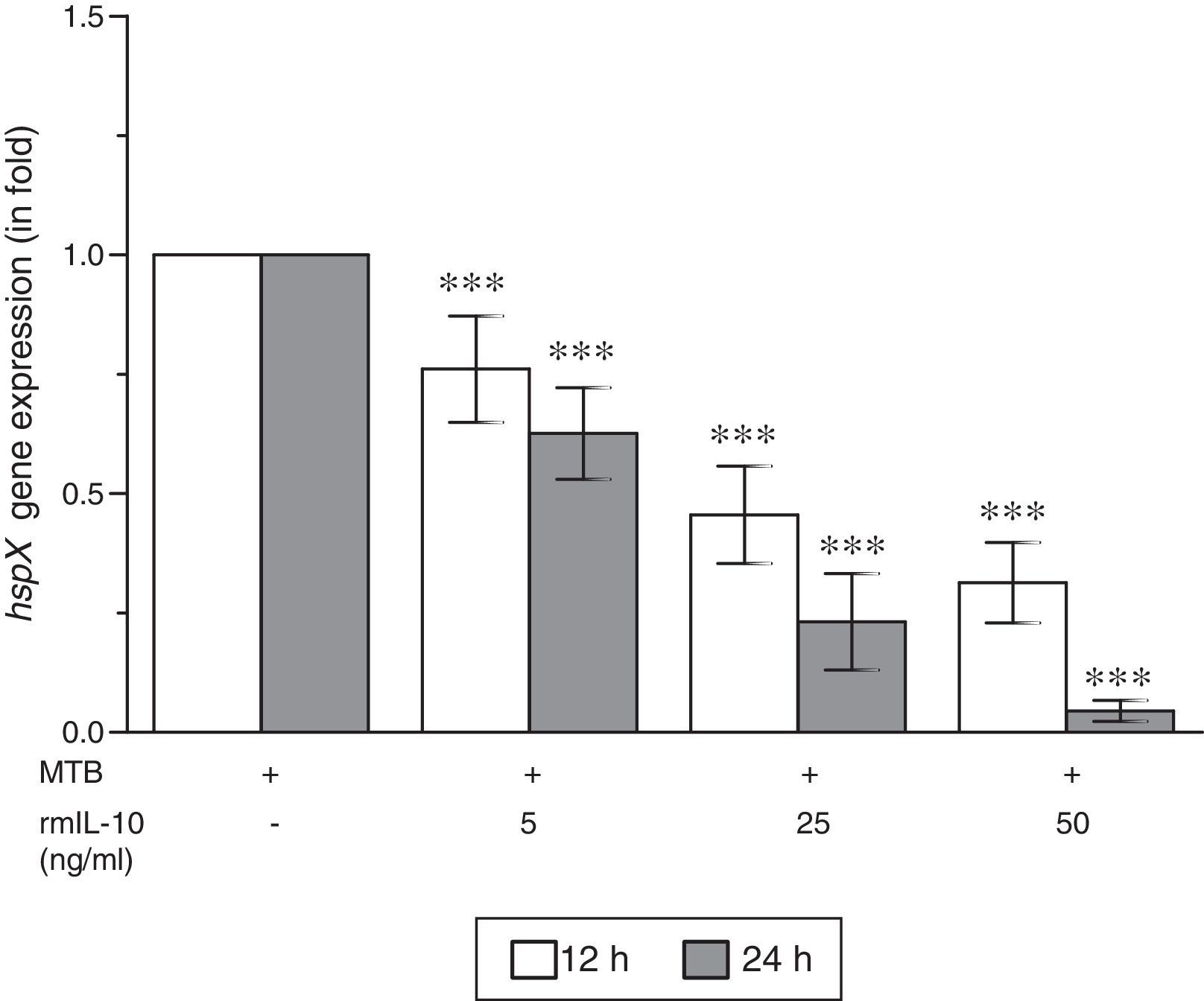
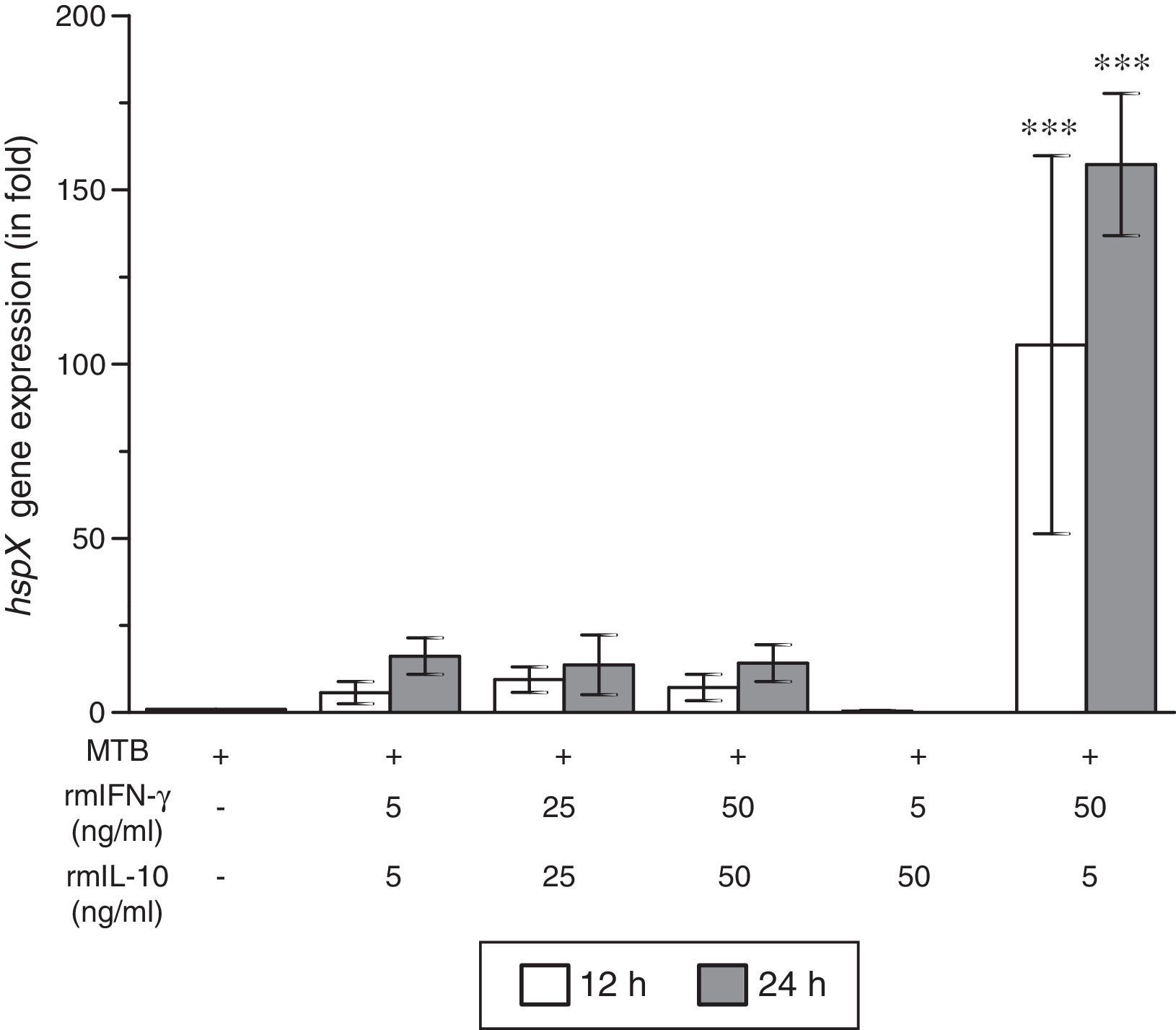
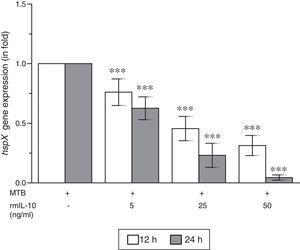
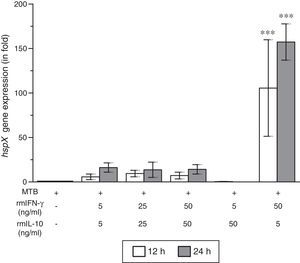
ResultsThe present study was carried out to see whether treatment of IFN-γ and IL-10 cytokines to intracellular MTB, engulfed by RAW264.7 cells, modulates the expression of its gene hspX encoding small heat shock protein 16.3. For this purpose, three treatment groups were made: (i) rmIFN-γ; (ii) rmIL-10; and (iii) rmIFN-γ and rmIL-10. Two time-points, i.e. 12 and 24h were used for each of the above groups. In the first group, RAW264.7 cells were infected with H37Rv strain of MTB at 1:15 MOI following treatment with three increasing concentrations (5, 25 and 50ng/ml) of rmIFN-γ. The data obtained from real-time RT-PCR analysis showed that treatment of intracellular MTB with aforesaid concentrations of rmIFN-γ resulted in significant up-regulation of hspX expression in concentration as well as in a time-dependent manner (Fig. 1). In the second group, the same concentrations, i.e. 5, 25 and 50ng/ml of rmIL-10 was used to treat the intracellular MTB at the same specified time-points and the obtained findings were quite different. Treatment of intracellular MTB with different concentrations of rmIL-10 caused significant down-regulation of hspX expression in concentration- and time-dependent fashion (Fig. 2). In the combination group (group 3) in which the MTB infected RAW264.7 cells were incubated with combined doses of rmIFN-γ and rmIL-10 for 12 and 24h, respectively, it was found that out of five combined doses, a dose consisting of 5ng/ml of rmIFN-γ and 50ng/ml of rmIL-10 down-regulates the expression of hspX at both time-points in comparison with untreated controls whereas a dose consisting of 50ng/ml of rmIFN-γ and 5ng/ml of rmIL-10 significantly up-regulates the induction of hspX in time-dependent manner. The rest of the three combined doses (rmIFN-γ, 5ng/ml+rmIL-10, 5ng/ml; rmIFN-γ, 25ng/ml+rmIL-10, 25ng/ml; and rmIFN-γ, 50ng/ml+rmIL-10, 50ng/ml) did not show any significant effect on the expression of hspX at both the experimental time-points (Fig. 3).
Effect of rmIFN-γ on the expression of gene hspX of MTB in RAW264.7 cells. The 2−ΔΔCT method was used to analyze the relative changes in expression of hspX in comparison with 16S rRNA. The values are the mean±S.D. of three independent experiments. *p<0.05, ***p<0.0001 between control and treated samples.
Effect of rmIL-10 on the expression of gene hspX of MTB in RAW264.7 cells. The 2−ΔΔCT method was used to analyze the relative changes in expression of hspX in comparison with 16S rRNA. The values are the mean±S.D. of three independent experiments. *p<0.05, ***p<0.0001 between control and treated samples.
Effect of rmIFN-γ and rmIL-10 on the expression of gene hspX of MTB in RAW264.7 cells. The 2−ΔΔCT method was used to analyze the relative changes in expression of hspX in comparison with 16S rRNA. The values are the mean±S.D. of three independent experiments. *p<0.05, ***p<0.0001 between control and treated samples.
Small heat shock proteins have been identified as vital biomolecules having a substantial role in the pathogenesis of MTB.2,20 MTB has two small heat shock proteins: Acr1 (α-crystallin related protein 1/sHSP16.3/HSP16.3/16kDa antigen/HspX2) and Acr2/HrpA.21 sHSP16.3 is one of the crucial proteins of MTB that facilitates survival of MTB in its host during prolonged periods of infection13 most probably by altering the expression of certain microRNAs.22 Expression of hspX has been found to be mainly regulated by a two-component response regulator dosR, dormancy survival regulator (previously known as devR, Rv3133c).23 The factors which trigger the induction of hspX are not fully known. However, it is suggested that hypoxia is a major stimulus behind the expression of this gene.2 Besides, NO,15 vitamin C,16 and ingestion of MTB by macrophage11 have also been reported to up-regulate the expression of hspX.
Cytokines are potent immunomodulators having pronounced effects on immunologic responses generated by a number of cell types in normal as well as in pathologic conditions. Although the role of IFN-γ and IL-10 in the pathogenesis of tuberculosis as well as in the intracellular survival of MTB24 is well known, their effect on the expression of survival associated gene hspX has not been previously reported.
Our study has demonstrated that three increasing concentrations (5, 25, and 50ng/ml) of rmIFN-γ significantly up-regulated the expression of hspX whereas similar concentrations of rmIL-10 (5, 25, and 50ng/ml) significantly down-regulated the hspX expression. This effect was found to be dependent on both the concentration of the stimulus and time. A contrasting pattern of hspX expression was observed when MTB infected macrophages were exposed to combinations of two different concentrations of rmIFN-γ and rmIL-10, i.e. low expression of hspX to a combination of low concentration of rmIFN-γ and high concentration of rmIL-10 versus high expression of hspX to a combination of high concentration of rmIFN-γ and low concentration of rmIL-10. The mechanisms of such cytokine mediated modulation of hspX expression are yet to be fully explored, which may be mediated through dormancy regulon dosR. An earlier study has reported that targeted disruption of this regulon results in the elimination of hspX expression.23
Growth retardation is a characteristic feature of mycobacterial dormancy.25 Studies carried out earlier with mutant strains showed that over-expression of hspX gene slows the growth of both MTB and Mycobacterium smegmatis2 and probably pushes the bacteria to enter dormant state.13 As hspX appears to play a central role in dormancy of MTB and potent anti-tuberculosis drugs do not effectively act on dormant organisms,26 rIL-10 driven down-regulation of hspX expression, in the presence of low concentration of rIFN-γ, would possibly decrease the chances of developing dormancy and increase the intracellular growth of MTB. Thus, it would be a better opportunity to anti-tuberculosis drugs to act on multiplying MTB. Such intervention could also be of help in prevention/elimination of persisters and in shortening of therapy. This may be too optimistic to rely only on one gene or biomarker like hspX as a number of genes are known to be associated with survival of MTB inside the host.1,27–29 At this moment, this is essentially a hypothesis which needs to be validated first in human cell lines and subsequently in experimental animal models mimicking the latent infection using clinical isolates of MTB. If successful, this may then be validated in human subjects by following proper approved and well planned protocols.
ConclusionThis study demonstrates that rmIL-10 down-regulates the expression of gene hspX of MTB in murine macrophage. Further studies involving other experimental models are warranted to validate the findings of this study.
Conflicts of interestThe authors declare no conflicts of interest.
The financial support from Department of Science and Technology, New Delhi in the form of Project No. SR/SO/HS-66/2007 is duly acknowledged. Authors are highly thankful to Director, International Center for Genetic Engineering and Biotechnology, New Delhi for providing essential research facilities; Arvind Pandey, Director, National Institute of Medical Statistics, New Delhi for statistical analysis; Santosh Kumar and Zaved Siddiqui, International Center for Genetic Engineering and Biotechnology, New Delhi for their technical support.