There are a lot of disagreements in the studies on hepatitis B virus (HBV) DNA polymerase mutation rate associated with nucleos(t)ide analogues (NAs) in treatment-naive chronic hepatitis B (CHB) patients. This is the first study aimed to investigate the prevalence of spontaneous HBV resistance mutations in Central China.
MethodsThis study included treatment-naive patients with CHB from June 2012 to May 2015 receiving care at the Institute of Liver Disease in Central China. All patients completed a questionnaire covering different aspects, such as family medical history, course of liver disease, medication history, alcohol use, among others. Mutations in HBV DNA polymerase associated with NAs resistance were detected using INNO-LiPA assay.
Results269 patients were infected with HBV genotype B (81.4%), C (17.9%), and both B and C (0.7%). Mutations in HBV DNA polymerase were detected in 24 patients (8.9%) including rtM204I/V (n=6), rtN236T (n=5), rtM250V (n=2), rtL180M (n=2), rtT184G (n=1), rtM207I (n=1), rtS202I (n=1), rtM204V/I & rtL180M (n=5), and rtM204I & rtM250V (n=1).
ConclusionSpontaneous HBV resistance mutations in HBV DNA polymerase were found in treatment-naive patients with CHB in Central China. These findings suggest that we should analyze HBV DNA polymerase resistance mutation associated with NAs before giving antiviral therapy such as lamivudine (LAM), adefovir (ADV), and telbivudine (LdT).
Infection with hepatitis B virus (HBV) can cause severe diseases such as liver cirrhosis and hepatocellular carcinoma (HCC), with approximately one million deaths annually around the world.1 Nucleos(t)ide analogues (NAs), including lamivudine (LAM), adefovir (ADV), telbivudine (LdT), entecavir (ETV), and tenofovir (TDF) were recommended by international guidelines for suppressing HBV replication, and have been shown to decrease the rate of complications.2–4
Despite high drug efficacy for inhibiting HBV, one of the largest obstacles with long-term NAs therapy is the development of viral resistance. LAM, used widely as the first NA, has the highest viral resistance rate of 70% after five years of therapy.5 ADV and LdT are superior to LAM for their lower prevalence of viral resistance, while ETV and TDF effectively suppress HBV DNA replication with minimum drug resistance.6 NAs treatment failure has been linked to mutations at the reverse transcriptase (rt) region of the HBV DNA polymerase gene, and mutations at individual codons in the HBV polymerase gene conferring resistance to different NAs. For example, rtM204V/I pathway is responsible for the resistance to L-NAs, such as LAM, LdT, clevudine (CLD), and ETV. Also, rtN236T pathway could confer ADV and TDF resistance, and rtA181T/V pathway could affect LAM, ADV, LdT, and TDF efficacy.5,7,8
It has been found that natural HBV reverse transcriptase mutations exist even in treatment-naive patients with CHB from Europe, Asia, and the Middle East, but the prevalence vary from 0% to 57%.9–13 This wide range might be caused by different study designs, regions, ethnicities, mutation detection methods, sample sizes, and so on. Using a systematic review and meta-analysis, Zhang et al.13 found an 8.0% prevalence of total and primary mutations among untreated CHB patients in China, and that Southern China had a little higher pooled total mutation rate of 8.22% than Northern China 7.55%. Therefore, it is important to understand the true prevalence of HBV DNA polymerase mutation analysis among treatment-naive patients with CHB in different areas of China, because most of them will be treated with NAs at first. To our knowledge, there has been no study in patients of Central China. The purpose of this study is to determine the prevalence and clinical characteristics of spontaneous mutations in Central Chinese cohort, and focus on clinically significant HBV DNA polymerase associated with NAs resistance mutations (rtM250V, rtN236T, rtM204V/I, rtS202I, rtA181T/V, rtT184G, rtL180M, and rtM/V207I) by using the sensitive INNO-LiPA assay.
Patients and methodsPatientsWe performed a prospectively analysis of 269 patients with CHB of the Institute of Liver Disease, Hubei Provincial Hospital of Traditional Chinese Medicine, Wuhan who had never received any anti-HBV treatment with NAs or interferon, between June 2012 and May 2015. Patients were included according to the following criteria – age 18 years or older, HBV DNA >100IU/mL, positive hepatitis B surface antigen, history of HBV infection for more than 6 months. Patients co-infected with HCV, HDV, or HIV were excluded. The diagnosis of hepatitis was established by needle biopsy. Liver biopsies were performed under ultrasound guiding and histological grade (G) and stage (S) were evaluated. We collected patients of chronic HBV infection and divided them into four phases (immune tolerant phase, immune clearance phase, low or non-replicative phase, reactivation phase) by the standard of the Guidelines for Chronic Hepatitis B in China, version 2010. The informed consent was obtained from each patient enrolled in the study. The study protocol conformed to the ethical guidelines of the Declaration of Helsinki and was approved by the ethics committee of Hubei Provincial Hospital of Traditional Chinese Medicine.
Patient questionnairesQuestionnaires, including questions about family medical history, HBV infection and treatment history, other disease history, ethnicity, and drug and alcohol history were completed by all patients.
General laboratory testsMarkers of HBV infection were measured using the commercial kit of electro-chemiluminescence assay (Roche Diagnostics, Mannheim, Germany). HBV DNA was quantified using the Amplicon monitor assay (ABI, California, USA). ALT, AST, and GGT were determined using the commercial kit of ultraviolet absorption spectrophotometry assay (Roche Diagnostics, Mannheim, Germany).
HBV polymerase gene mutation and genotype by INNO-LiPAHBV DNA polymerase mutations and genotypes were measured using the INNO-LiPA assay kit (Shenzhen, China). This assay detected mutations at codons 180, 181, 184, 202, 204, 207, 236, 250 of the HBV DNA polymerase gene, with a reported detection lower bound of 5% of the circulating viral load.
Statistical analysisResults were reported as mean and standard deviation and/or median and range for continuous variables and percentages for categorical variables. The categorical data were compared between groups using the chi-square test or Fisher's exact test, and the normally distributed continuous variables were analyzed by the Student's t-test. The nonparametric variables were analyzed by the Mann–Whitney U-test. All p-values were two-tailed, and p<0.05 was considered statistically significant. Data analysis was performed using the Statistical Package for Social Science version 19.0 (SPSS, Chicago, USA).
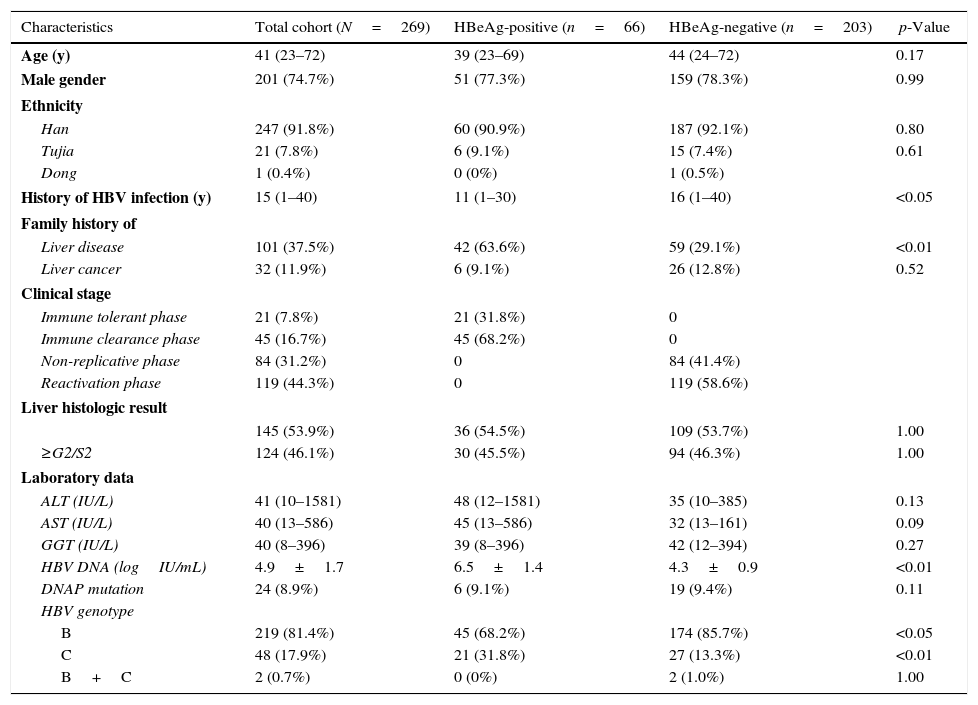
ResultsPatient characteristicsThe characteristics of participating patients at baseline are reported in Table 1. Of the 269 patients, 74.7% (201/269) were male with a median age of 41 years. At baseline, 75.5% (203/269) of the patients were HBeAg negative, and the HBV DNA load was 4.9logIU/mL. Furthermore, 81.4% (219/269), 17.9% (48/269) and 0.7% (2/269) patients were infected with HBV genotype B, C, and B & C, respectively.
Characteristics of patients according to the status of HBeAg.
| Characteristics | Total cohort (N=269) | HBeAg-positive (n=66) | HBeAg-negative (n=203) | p-Value |
|---|---|---|---|---|
| Age (y) | 41 (23–72) | 39 (23–69) | 44 (24–72) | 0.17 |
| Male gender | 201 (74.7%) | 51 (77.3%) | 159 (78.3%) | 0.99 |
| Ethnicity | ||||
| Han | 247 (91.8%) | 60 (90.9%) | 187 (92.1%) | 0.80 |
| Tujia | 21 (7.8%) | 6 (9.1%) | 15 (7.4%) | 0.61 |
| Dong | 1 (0.4%) | 0 (0%) | 1 (0.5%) | |
| History of HBV infection (y) | 15 (1–40) | 11 (1–30) | 16 (1–40) | <0.05 |
| Family history of | ||||
| Liver disease | 101 (37.5%) | 42 (63.6%) | 59 (29.1%) | <0.01 |
| Liver cancer | 32 (11.9%) | 6 (9.1%) | 26 (12.8%) | 0.52 |
| Clinical stage | ||||
| Immune tolerant phase | 21 (7.8%) | 21 (31.8%) | 0 | |
| Immune clearance phase | 45 (16.7%) | 45 (68.2%) | 0 | |
| Non-replicative phase | 84 (31.2%) | 0 | 84 (41.4%) | |
| Reactivation phase | 119 (44.3%) | 0 | 119 (58.6%) | |
| Liver histologic result | ||||
| 145 (53.9%) | 36 (54.5%) | 109 (53.7%) | 1.00 | |
| ≥G2/S2 | 124 (46.1%) | 30 (45.5%) | 94 (46.3%) | 1.00 |
| Laboratory data | ||||
| ALT (IU/L) | 41 (10–1581) | 48 (12–1581) | 35 (10–385) | 0.13 |
| AST (IU/L) | 40 (13–586) | 45 (13–586) | 32 (13–161) | 0.09 |
| GGT (IU/L) | 40 (8–396) | 39 (8–396) | 42 (12–394) | 0.27 |
| HBV DNA (logIU/mL) | 4.9±1.7 | 6.5±1.4 | 4.3±0.9 | <0.01 |
| DNAP mutation | 24 (8.9%) | 6 (9.1%) | 19 (9.4%) | 0.11 |
| HBV genotype | ||||
| B | 219 (81.4%) | 45 (68.2%) | 174 (85.7%) | <0.05 |
| C | 48 (17.9%) | 21 (31.8%) | 27 (13.3%) | <0.01 |
| B+C | 2 (0.7%) | 0 (0%) | 2 (1.0%) | 1.00 |
Note: Results are presented as proportion (%), mean±standard deviation, or median (range).
ALT, alanine aminotransferase; AST, aspartate aminotransferase; GGT, γ-glutamyl-transferase; DNAP, HBV DNA polymerase.
HBeAg-positive patients had a shorter median time of HBV infection (p<0.05), and were significantly more likely to report a family history of liver disease (p<0.01), compared with HBeAg-negative patients. HBeAg-positive patients had significantly higher baseline HBV DNA loads than HBeAg-negative (p<0.01). HBeAg-positive patients were more likely to have HBV genotype C infection than HBeAg-negative (p<0.01), but, contrary to the latter, had higher prevalence rate of HBV genotype B than the former (p<0.05).
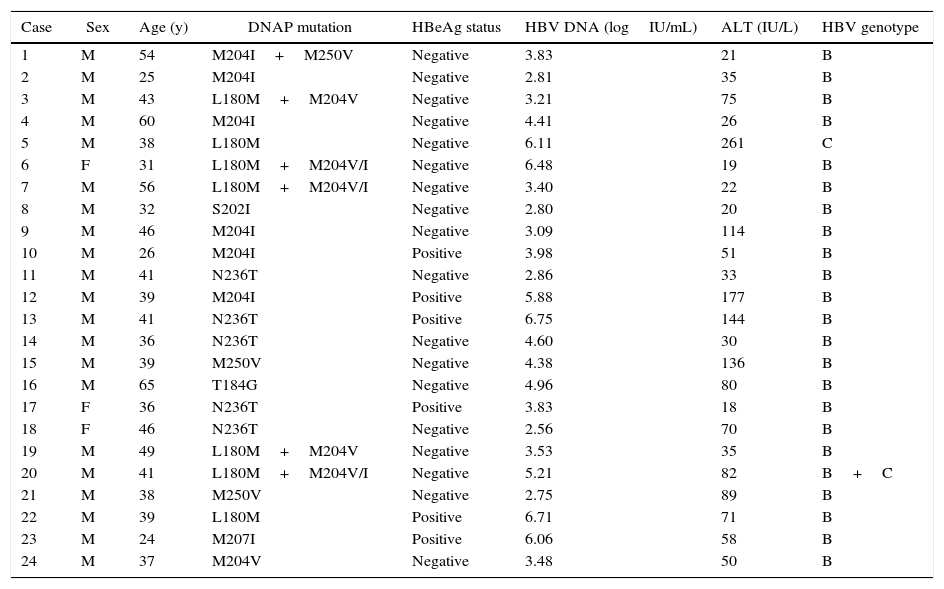
Prevalence of HBV DNA polymerase mutationsTable 2 lists the characteristics of patients with HBV DNA polymerase mutations based on INNO-LiPA assay. HBV DNA polymerase mutations were detected in 24 of 269 total patients (8.9%), including rtM204I/V (n=6), rtN236T (n=5), rtM250V (n=2), rtL180M (n=2), rtT184G (n=1), rtM207I (n=1), rtS202I (n=1), rtM204V/I & rtL180M (n=5), and rtM204I & rtM250V (n=1). Among all the mutations, rtM204V/I ranked first with the highest prevalence of 4.5% (12/269), which is larger than the prevalence rate of rtL180M, rtN236T, rtM250V, rtT184G, rtS202I, and rtV207I (2.6%, 1.9%, 1.1%, 0.4%, 0.4%, and 0.4%, respectively). Of 24 patients with HBV DNA polymerase mutations, 87.5% (21/24) were male, and the median age was 39 years. In addition, the mean HBV DNA load was 4.34logIU/mL, and 25.0% (6/24) of the patients were HBeAg positive with HBV genotype B. Only one HBV C-genotype, and one HBV C+B-genotype were detected among the 24 treatment-naive CHB patients for HBV DNA polymerase mutations.
Characteristics of patients with HBV DNA polymerase mutations.
| Case | Sex | Age (y) | DNAP mutation | HBeAg status | HBV DNA (logIU/mL) | ALT (IU/L) | HBV genotype |
|---|---|---|---|---|---|---|---|
| 1 | M | 54 | M204I+M250V | Negative | 3.83 | 21 | B |
| 2 | M | 25 | M204I | Negative | 2.81 | 35 | B |
| 3 | M | 43 | L180M+M204V | Negative | 3.21 | 75 | B |
| 4 | M | 60 | M204I | Negative | 4.41 | 26 | B |
| 5 | M | 38 | L180M | Negative | 6.11 | 261 | C |
| 6 | F | 31 | L180M+M204V/I | Negative | 6.48 | 19 | B |
| 7 | M | 56 | L180M+M204V/I | Negative | 3.40 | 22 | B |
| 8 | M | 32 | S202I | Negative | 2.80 | 20 | B |
| 9 | M | 46 | M204I | Negative | 3.09 | 114 | B |
| 10 | M | 26 | M204I | Positive | 3.98 | 51 | B |
| 11 | M | 41 | N236T | Negative | 2.86 | 33 | B |
| 12 | M | 39 | M204I | Positive | 5.88 | 177 | B |
| 13 | M | 41 | N236T | Positive | 6.75 | 144 | B |
| 14 | M | 36 | N236T | Negative | 4.60 | 30 | B |
| 15 | M | 39 | M250V | Negative | 4.38 | 136 | B |
| 16 | M | 65 | T184G | Negative | 4.96 | 80 | B |
| 17 | F | 36 | N236T | Positive | 3.83 | 18 | B |
| 18 | F | 46 | N236T | Negative | 2.56 | 70 | B |
| 19 | M | 49 | L180M+M204V | Negative | 3.53 | 35 | B |
| 20 | M | 41 | L180M+M204V/I | Negative | 5.21 | 82 | B+C |
| 21 | M | 38 | M250V | Negative | 2.75 | 89 | B |
| 22 | M | 39 | L180M | Positive | 6.71 | 71 | B |
| 23 | M | 24 | M207I | Positive | 6.06 | 58 | B |
| 24 | M | 37 | M204V | Negative | 3.48 | 50 | B |
Note: M, male; F, female; ALT, alanine aminotransferase; DNAP, HBV DNA polymerase.
The occurrence of viral resistance to NAs is closely related with amino acid substitutions in HBV reverse transcriptase, defined as primary and secondary drug resistance mutations. There are currently eight codons associated with primary drug resistance to NAs: rtI169T, rtA181T/V, rtT184S/C/G/A, rtA194T, rtS202G/C/I, rtM204V/I, rtN236T, and rtM250V; and three codons associated with secondary resistance to NAs: rtL80I, rtV173L and rt180M.7 The primary mutations refer to the amino acid changes that result in reduced susceptibility to an antiviral agent. The secondary mutations cause amino acid substitutions that restore replication function defects resulting from primary resistance and may give rise to a low level of reduced susceptibility.14,15
Currently, mutations associated with HBV drug resistance are reported to occur in treatment-naive patients with CHB for any oral anti-HBV therapy.16–29 This might due to both HBV viral factors and host factors. First, these mutations occur naturally, because HBV has a high replication rate, with an estimated production of about 1012virions/day, and a high mutational rate of 10−5 to 10−4 substitutions/base/cycle. Moreover, the HBV reverse transcriptase does not have a proofreading function to repair incorrectly incorporated nucleotides.30 Second, a treatment-naive patient might be infected by a mutant virus derived from a NAs treated patient. Third, the patient could be unaware of receiving some treatment with direct anti-HBV activity.
This study has confirmed the existence of HBV DNA polymerase resistance mutations for NAs in 24 (8.9%) of 269 treatment-naive patients, composed of five primary drug resistance mutations (rtT184G, rtS202I, rtM204I/V, rtN236T, and rtM250V), one secondary resistance mutation (rtL180M), and one codon mutation that might be baseline polymorphisms without any clinical significance (rtV207I). Among the 24 mutations, rtM204V/I was the prevailing type with a higher prevalence of 4.5% (12/269), compared with other codons with prevalence as low as 2.6% (7/269) of rtL180M, 1.9% (5/269) of rtN236T, 1.1% (3/269) of rtM250V, 0.4% (1/269) of rtT184G, 0.4% (1/269) of rtS202I, and 0.4% (1/269) of rtV207I. This means up to 6.5% treatment-naive patients with CHB would encounter LAM or LdT or ADV or ETV primary treatment failure initially, because rtM204V/I (YMDD/YIDD) and rtL180M linked to LAM and ETV resistance, rtM204I accounts for LdT resistance, and rtN236T could confer ADV resistance. Meanwhile, 2.2% (6/269) are multiple resistance mutations including 1.9% (5/269) rtM204V/I & rtL180M, and 0.4% (1/269) rtM204V/I & rtM250V. These treatment-naive patients with CHB would encounter ETV primary treatment failure initially, because the combination of rtM204V/I, rtL180M and one of the following mutations – rtM184S/C/G/A, rtS202G/I/C, or rtM250V – would lead to ETV resistance.7,8,16
To date, the actual prevalence of HBV DNA polymerase mutations in treatment-naive patients with CHB is unknown, and the reported prevalence varied from 0% to 57%.9,12,16–29 Such a wide range is likely due to differences in study design, study population and geography, study method, study sample size, and whether study patients were treatment-naive. Previous studies on the prevalence of drug resistance mutations in treatment-naive patients often used less sensitive methods or included rather small series of patients.9,19 Now a new sensitive ultra-deep pyrosequencing assay is able to detect minor viral quasispecies at a level under 1%.21 A systematic review and meta-analysis from Zhang et al.13 studied a total of 70 articles including 8156 treatment-naive patients with CHB in China, 21 articles of which were from Northern China and 49 from Southern China. They found that Southern China had a little higher pooled total mutation rate of 8.22% than Northern China (7.55%), as well as higher primary mutation rate of 8.12% versus 7.13%. The HBV variants known to confer resistance against NAs were detected and have a frequency of mutations quite comparable to what we found. In contrast, Nishijima et al.21 reported a relatively high mutation rate of 35.7% in 14 untreated patients with CHB in Japan using ultra-deep pyrosequencing. The difference between their studies and our results could be explained by a smaller sample size as well as the different method they used since ultra-deep pyrosequencing analysis is much more sensitive than INNO LiPA. Contrary to the above reports, several studies from Europe,9 North America19 and Latin America,28 reported that less than 5% primary mutation rate associated to NAs were found in treatment-naive patients. They used direct sequencing assay that could potentially underestimate the true prevalence because of its limited ability to reliably detect viral subspecies below 20%–25% of the circulating viral load.21 Meanwhile, the cohort of patients was predominantly infected with HBV genotypes A, D and C which were not common genotypes in China, especially Central China.
This study has the following advantages. Firstly, to our knowledge, this is the first comprehensive research about resistance mutations of the HBV polymerase in Central China, while all prior studies only focused on a few provinces and municipalities of Northern, Southern, and Eastern China such as Beijing, Shanghai, and Jiangsu.10,17,25,26,29 Secondly, all 269 patients have been required to finish a detailed questionnaire including questions about whether they accepted prior NA therapy, while the majority of other studies collected and analyzed serum for DNA polymerase mutations but did not validate whether the patients received treatment for NAs. Thirdly, we focused on mutations at the reverse transcriptase region of the HBV DNA polymerase gene that has been linked to different NAs treatment failure, while most researches merely focused on rtM204V/I, since LAM had the highest prevalence of resistance and YMDD mutation was extensively studied. Other primary mutations and secondary mutations associated with resistance to NAs did not receive enough attention. Lastly, the INNO-LiPA assay in our study is a sensitive and specific assay that can detect subspecies at 5% of the circulating viral population, while the direct sequencing assay is limited by its lower sensitivity that detects mutants only if they consist of greater than 20%–25% of the circulating viral load.12
In conclusion, the prevalence of HBV DNA polymerase mutations associated with NAs resistance was found to be up to 8.9% of our 269 treatment-naive patients with CHB in Central China. The LAM-associated rtM204V/I mutation occurred more frequently than other primary mutations. In China, LAM, ADV, and LdT are commonly used alone or together, ETV and TDF are seldom used. These findings suggest that HBV DNA polymerase associated with NAs resistance mutation should be analyzed before initiating antiviral therapy such as LAM, ADV, and LdT for the patients with CHB.
Conflicts of interestThe authors declare no conflicts of interest.
This work was supported by the special funds for State Administration of Traditional Chinese Medicine of the People's Republic of China (No. JDZX2012051).